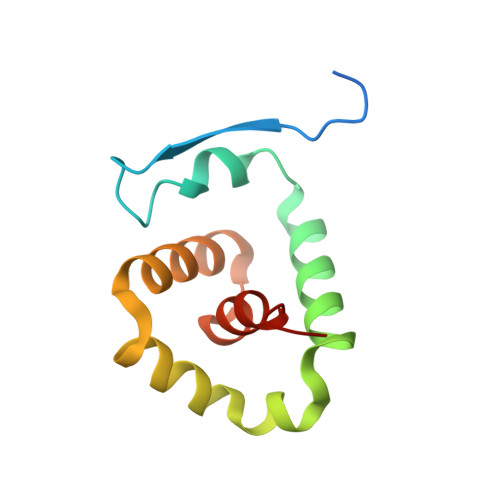
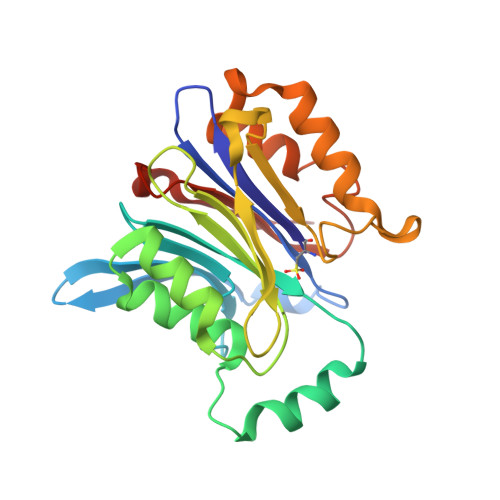
Molecular mechanism of activation of the immunoregulatory amidase NAAA.
Gorelik, A., Gebai, A., Illes, K., Piomelli, D., Nagar, B.(2018) Proc Natl Acad Sci U S A 115: E10032-E10040
- PubMed: 30301806
- DOI: https://doi.org/10.1073/pnas.1811759115
- Primary Citation Related Structures:
6DXW, 6DXX, 6DXY, 6DXZ, 6DY0, 6DY1, 6DY2, 6DY3 - PubMed Abstract:
Palmitoylethanolamide is a bioactive lipid that strongly alleviates pain and inflammation in animal models and in humans. Its signaling activity is terminated through degradation by N -acylethanolamine acid amidase (NAAA), a cysteine hydrolase expressed at high levels in immune cells. Pharmacological inhibitors of NAAA activity exert profound analgesic and antiinflammatory effects in rodent models, pointing to this protein as a potential target for therapeutic drug discovery. To facilitate these efforts and to better understand the molecular mechanism of action of NAAA, we determined crystal structures of this enzyme in various activation states and in complex with several ligands, including both a covalent and a reversible inhibitor. Self-proteolysis exposes the otherwise buried active site of NAAA to allow catalysis. Formation of a stable substrate- or inhibitor-binding site appears to be conformationally coupled to the interaction of a pair of hydrophobic helices in the enzyme with lipid membranes, resulting in the creation of a linear hydrophobic cavity near the active site that accommodates the ligand's acyl chain.
- Department of Biochemistry, McGill University, Montreal, H3G0B1, Canada.
Organizational Affiliation: