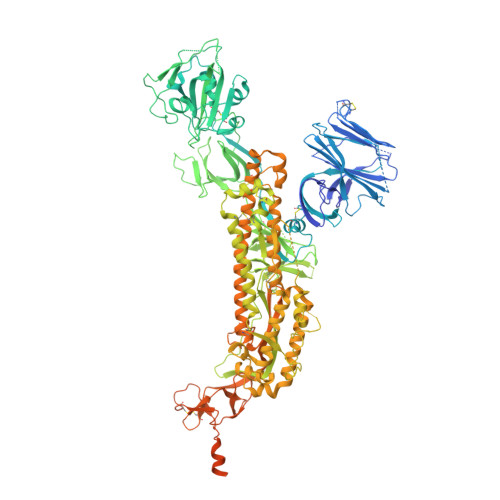
Stabilizing the closed SARS-CoV-2 spike trimer.
Juraszek, J., Rutten, L., Blokland, S., Bouchier, P., Voorzaat, R., Ritschel, T., Bakkers, M.J.G., Renault, L.L.R., Langedijk, J.P.M.(2021) Nat Commun 12: 244-244
- PubMed: 33431842 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-20321-x
- Primary Citation Related Structures:
7A4N, 7AD1 - PubMed Abstract:
The trimeric spike (S) protein of SARS-CoV-2 is the primary focus of most vaccine design and development efforts. Due to intrinsic instability typical of class I fusion proteins, S tends to prematurely refold to the post-fusion conformation, compromising immunogenic properties and prefusion trimer yields. To support ongoing vaccine development efforts, we report the structure-based design of soluble S trimers with increased yields and stabilities, based on introduction of single point mutations and disulfide-bridges. We identify regions critical for stability: the heptad repeat region 1, the SD1 domain and position 614 in SD2. We combine a minimal selection of mostly interprotomeric mutations to create a stable S-closed variant with a 6.4-fold higher expression than the parental construct while no longer containing a heterologous trimerization domain. The cryo-EM structure reveals a correctly folded, predominantly closed pre-fusion conformation. Highly stable and well producing S protein and the increased understanding of S protein structure will support vaccine development and serological diagnostics.
- Janssen Vaccines & Prevention B.V., Archimedesweg 4-6, Leiden, The Netherlands.
Organizational Affiliation: