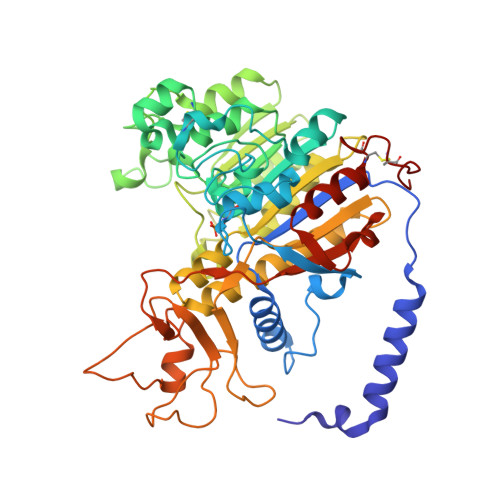
Structural Studies of Human Placental Alkaline Phosphatase in Complex with Functional Ligands.
Llinas, P., Stura, E.A., Menez, A., Kiss, Z., Stigbrand, T., Millan, J.L., Le Du, M.H.(2005) J Mol Biology 350: 441-451
- PubMed: 15946677 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.04.068
- Primary Citation Related Structures:
1ZEB, 1ZED, 1ZEF - PubMed Abstract:
The activity of human placental alkaline phosphatase (PLAP) is downregulated by a number of effectors such as l-phenylalanine, an uncompetitive inhibitor, 5'-AMP, an antagonist of the effects of PLAP on fibroblast proliferation and by p-nitrophenyl-phosphonate (PNPPate), a non-hydrolysable substrate analogue. For the first two, such regulation may be linked to its biological function that requires a reduced and better-regulated hydrolytic rate. To understand how such disparate ligands are able to inhibit the enzyme, we solved the structure of the complexes at 1.6A, 1.9A and 1.9A resolution, respectively. These crystal structures are the first of an alkaline phosphatase in complex with organic inhibitors. Of the three inhibitors, only l-Phe and PNPPate bind at the active site hydrophobic pocket, providing structural data on the uncompetitive inhibition process. In contrast, all three ligands interact at a remote peripheral site located 28A from the active site. In order to extend these observations to the other members of the human alkaline phosphatase family, we have modelled the structures of the other human isozymes and compared them to PLAP. This comparison highlights the crucial role played by position 429 at the active site in the modulation of the catalytic process, and suggests that the peripheral binding site may be involved in the functional specialization of the PLAP isozyme.
- Laboratoire de Structure des Protéines, Département d'Ingénierie et d'Etude des Protéines, Commissariat à l'Energie Atomique, 91191 Gif sur Yvette, France.
Organizational Affiliation: