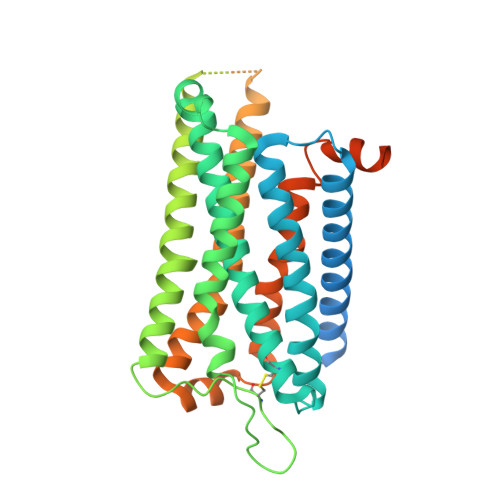
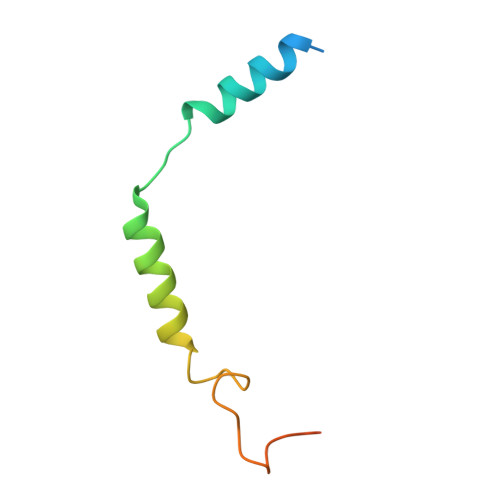
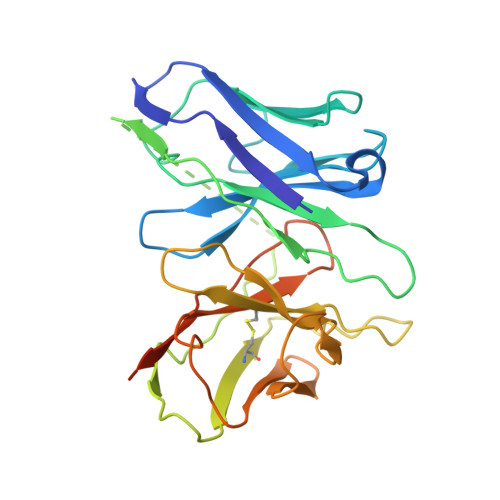
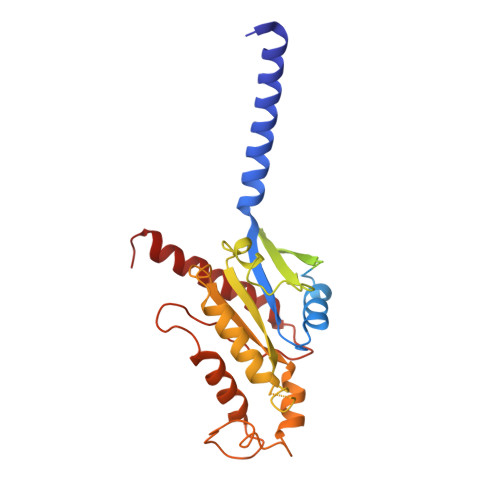
Structural insights into human brain-gut peptide cholecystokinin receptors.
Ding, Y., Zhang, H., Liao, Y.Y., Chen, L.N., Ji, S.Y., Qin, J., Mao, C., Shen, D.D., Lin, L., Wang, H., Zhang, Y., Li, X.M.(2022) Cell Discov 8: 55-55
- PubMed: 35672283 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-022-00420-3
- Primary Citation Related Structures:
7XOU, 7XOV, 7XOW, 8IA7 - PubMed Abstract:
The intestinal hormone and neuromodulator cholecystokinin (CCK) receptors CCK1R and CCK2R act as a signaling hub in brain-gut axis, mediating digestion, emotion, and memory regulation. CCK receptors exhibit distinct preferences for ligands in different posttranslational modification (PTM) states. CCK1R couples to G s and G q , whereas CCK2R primarily couples to G q . Here we report the cryo-electron microscopy (cryo-EM) structures of CCK1R-G s signaling complexes liganded either by sulfated cholecystokinin octapeptide (CCK-8) or a CCK1R-selective small-molecule SR146131, and CCK2R-G q complexes stabilized by either sulfated CCK-8 or a CCK2R-selective ligand gastrin-17. Our structures reveal a location-conserved yet charge-distinct pocket discriminating the effects of ligand PTM states on receptor subtype preference, the unique pocket topology underlying selectivity of SR146131 and gastrin-17, the conformational changes in receptor activation, and key residues contributing to G protein subtype specificity, providing multiple structural templates for drug design targeting the brain-gut axis.
- Department of Neurobiology and Department of Neurology of Second Affiliated Hospital, Zhejiang University School of Medicine, Hangzhou, Zhejiang, China.
Organizational Affiliation: