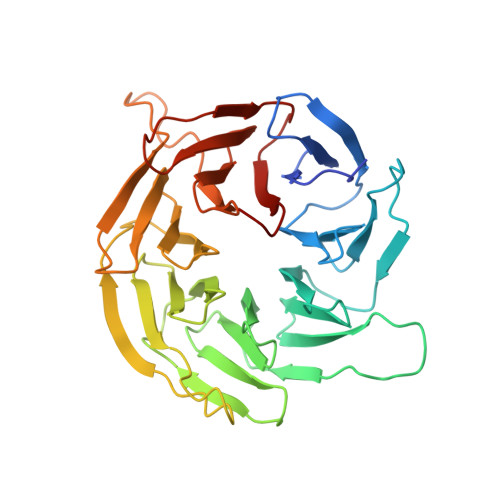
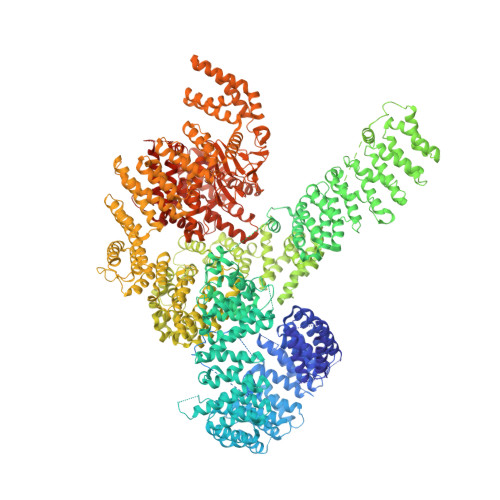
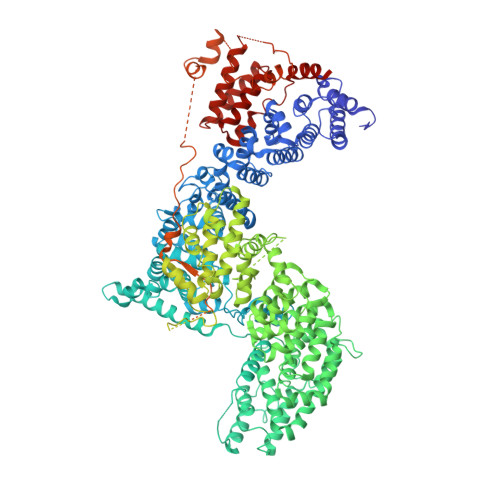
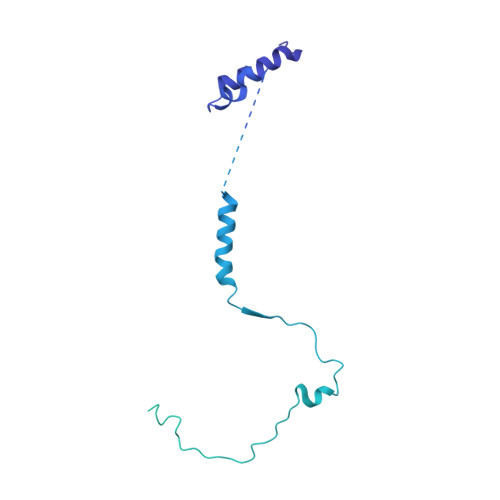
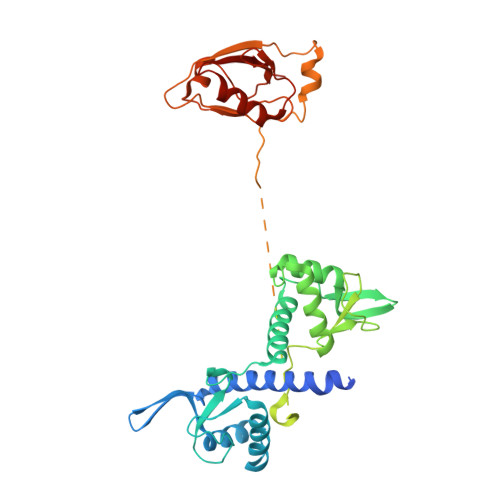
Regulation of human mTOR complexes by DEPTOR.
Walchli, M., Berneiser, K., Mangia, F., Imseng, S., Craigie, L.M., Stuttfeld, E., Hall, M.N., Maier, T.(2021) Elife 10
- PubMed: 34519268 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.70871
- Primary Citation Related Structures:
7PE7, 7PE8, 7PE9, 7PEA, 7PEB, 7PEC, 7PED - PubMed Abstract:
The vertebrate-specific DEP domain-containing mTOR interacting protein (DEPTOR), an oncoprotein or tumor suppressor, has important roles in metabolism, immunity, and cancer. It is the only protein that binds and regulates both complexes of mammalian target of rapamycin (mTOR), a central regulator of cell growth. Biochemical analysis and cryo-EM reconstructions of DEPTOR bound to human mTOR complex 1 (mTORC1) and mTORC2 reveal that both structured regions of DEPTOR, the PDZ domain and the DEP domain tandem (DEPt), are involved in mTOR interaction. The PDZ domain binds tightly with mildly activating effect, but then acts as an anchor for DEPt association that allosterically suppresses mTOR activation. The binding interfaces of the PDZ domain and DEPt also support further regulation by other signaling pathways. A separate, substrate-like mode of interaction for DEPTOR phosphorylation by mTOR complexes rationalizes inhibition of non-stimulated mTOR activity at higher DEPTOR concentrations. The multifaceted interplay between DEPTOR and mTOR provides a basis for understanding the divergent roles of DEPTOR in physiology and opens new routes for targeting the mTOR-DEPTOR interaction in disease.
- Biozentrum, University of Basel, Basel, Switzerland.
Organizational Affiliation: