A Spike-destructing human antibody effectively neutralizes Omicron-included SARS-CoV-2 variants with therapeutic efficacy.
Meng, L., Zha, J., Zhou, B., Cao, L., Jiang, C., Zhu, Y., Li, T., Lu, L., Zhang, J., Yang, H., Feng, J., Gu, Z., Tang, H., Jiang, L., Li, D., Lavillette, D., Zhang, X.(2023) PLoS Pathog 19: e1011085-e1011085
- PubMed: 36706160 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1011085
- Primary Citation Related Structures:
7WQV - PubMed Abstract:
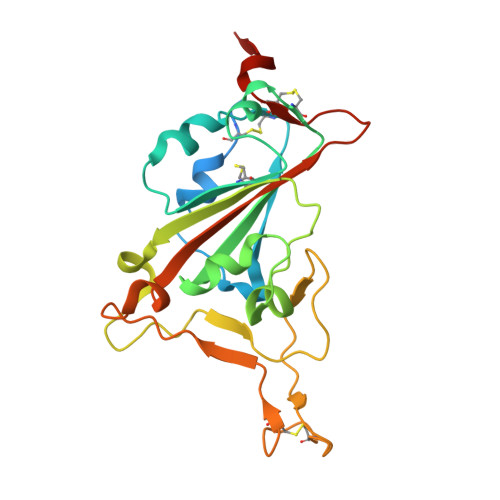
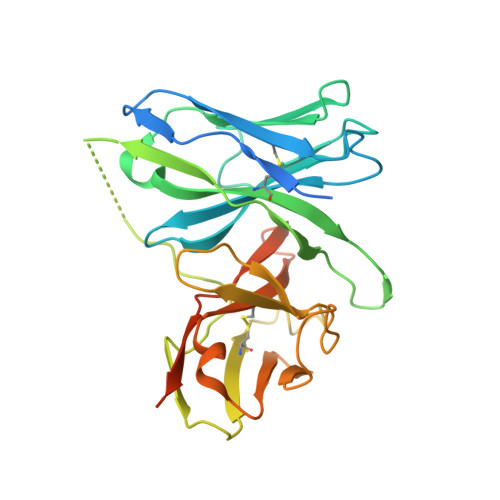
Neutralizing antibodies (nAbs) are important assets to fight COVID-19, but most existing nAbs lose the activities against Omicron subvariants. Here, we report a human monoclonal antibody (Ab08) isolated from a convalescent patient infected with the prototype strain (Wuhan-Hu-1). Ab08 binds to the receptor-binding domain (RBD) with pico-molar affinity (230 pM), effectively neutralizes SARS-CoV-2 and variants of concern (VOCs) including Alpha, Beta, Gamma, Mu, Omicron BA.1 and BA.2, and to a lesser extent for Delta and Omicron BA.4/BA.5 which bear the L452R mutation. Of medical importance, Ab08 shows therapeutic efficacy in SARS-CoV-2-infected hACE2 mice. X-ray crystallography of the Ab08-RBD complex reveals an antibody footprint largely in the β-strand core and away from the ACE2-binding motif. Negative staining electron-microscopy suggests a neutralizing mechanism through which Ab08 destructs the Spike trimer. Together, our work identifies a nAb with therapeutic potential for COVID-19.
- Key Laboratory of Molecular Virology & Immunology, Institut Pasteur of Shanghai, Chinese Academy of Sciences/University of Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: