Surface Ig variable domain glycosylation affects autoantigen binding and acts as threshold for human autoreactive B cell activation.
Kissel, T., Ge, C., Hafkenscheid, L., Kwekkeboom, J.C., Slot, L.M., Cavallari, M., He, Y., van Schie, K.A., Vergroesen, R.D., Kampstra, A.S.B., Reijm, S., Stoeken-Rijsbergen, G., Koeleman, C., Voortman, L.M., Heitman, L.H., Xu, B., Pruijn, G.J.M., Wuhrer, M., Rispens, T., Huizinga, T.W.J., Scherer, H.U., Reth, M., Holmdahl, R., Toes, R.E.M.(2022) Sci Adv 8: eabm1759-eabm1759
- PubMed: 35138894 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abm1759
- Primary Citation Related Structures:
6YXK, 6YXL, 6YXM - PubMed Abstract:
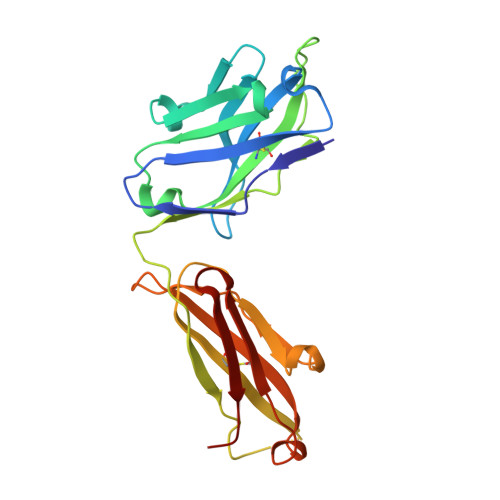
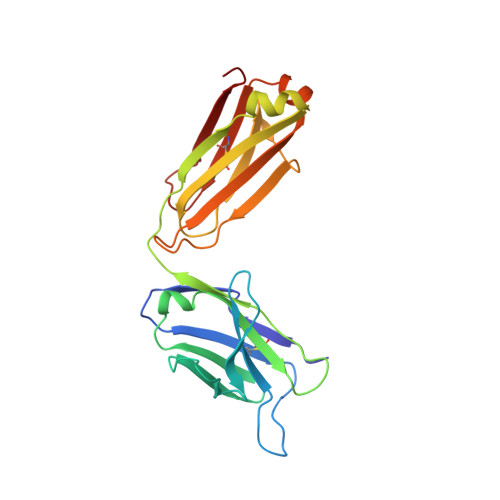
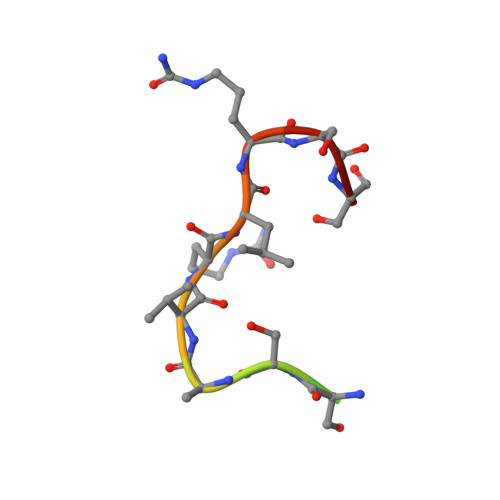
The hallmark autoantibodies in rheumatoid arthritis are characterized by variable domain glycans (VDGs). Their abundant occurrence results from the selective introduction of N-linked glycosylation sites during somatic hypermutation, and their presence is predictive for disease development. However, the functional consequences of VDGs on autoreactive B cells remain elusive. Combining crystallography, glycobiology, and functional B cell assays allowed us to dissect key characteristics of VDGs on human B cell biology. Crystal structures showed that VDGs are positioned in the vicinity of the antigen-binding pocket, and dynamic modeling combined with binding assays elucidated their impact on binding. We found that VDG-expressing B cell receptors stay longer on the B cell surface and that VDGs enhance B cell activation. These results provide a rationale on how the acquisition of VDGs might contribute to the breach of tolerance of autoreactive B cells in a major human autoimmune disease.
- Department of Rheumatology, Leiden University Medical Center, Leiden, Netherlands.
Organizational Affiliation: