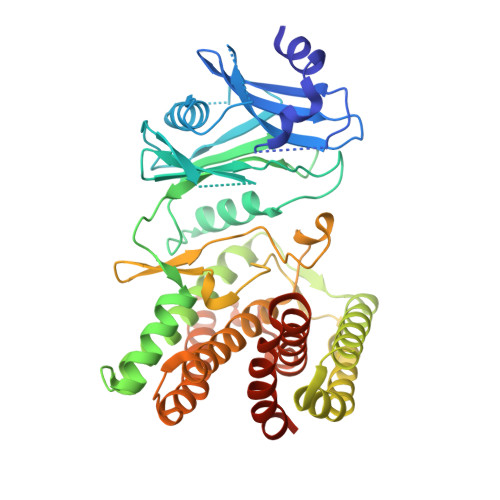
Molecular Fingerprints for a Novel Enzyme Family in Actinobacteria with Glucosamine Kinase Activity.
Manso, J.A., Nunes-Costa, D., Macedo-Ribeiro, S., Empadinhas, N., Pereira, P.J.B.(2019) mBio 10
- PubMed: 31088917
- DOI: https://doi.org/10.1128/mBio.00239-19
- Primary Citation Related Structures:
6HWJ, 6HWK, 6HWL - PubMed Abstract:
Actinobacteria have long been the main source of antibiotics, secondary metabolites with tightly controlled biosynthesis by environmental and physiological factors. Phosphorylation of exogenous glucosamine has been suggested as a mechanism for incorporation of this extracellular material into secondary metabolite biosynthesis, but experimental evidence of specific glucosamine kinases in Actinobacteria is lacking. Here, we present the molecular fingerprints for the identification of a unique family of actinobacterial glucosamine kinases. Structural and biochemical studies on a distinctive kinase from the soil bacterium Streptacidiphilus jiangxiensis unveiled its preference for glucosamine and provided structural evidence of a phosphoryl transfer to this substrate. Conservation of glucosamine-contacting residues across a large number of uncharacterized actinobacterial proteins unveiled a specific glucosamine binding sequence motif. This family of kinases and their genetic context may represent the missing link for the incorporation of environmental glucosamine into the antibiotic biosynthesis pathways in Actinobacteria and can be explored to enhance antibiotic production. IMPORTANCE The discovery of novel enzymes involved in antibiotic biosynthesis pathways is currently a topic of utmost importance. The high levels of antibiotic resistance detected worldwide threaten our ability to combat infections and other 20th-century medical achievements, namely, organ transplantation or cancer chemotherapy. We have identified and characterized a unique family of enzymes capable of phosphorylating glucosamine to glucosamine-6-phosphate, a crucial molecule directly involved in the activation of antibiotic production pathways in Actinobacteria , nature's main source of antimicrobials. The consensus sequence identified for these glucosamine kinases will help establish a molecular fingerprint to reveal yet-uncharacterized sequences in antibiotic producers, which should have an important impact in biotechnological and biomedical applications, including the enhancement and optimization of antibiotic production.
- IBMC-Instituto de Biologia Molecular e Celular, Universidade do Porto, Porto, Portugal.
Organizational Affiliation: