High-Resolution Structure of ClpC1-Rufomycin and Ligand Binding Studies Provide a Framework to Design and Optimize Anti-Tuberculosis Leads.
Wolf, N.M., Lee, H., Choules, M.P., Pauli, G.F., Phansalkar, R., Anderson, J.R., Gao, W., Ren, J., Santarsiero, B.D., Lee, H., Cheng, J., Jin, Y.Y., Ho, N.A., Duc, N.M., Suh, J.W., Abad-Zapatero, C., Cho, S.(2019) ACS Infect Dis 5: 829-840
- PubMed: 30990022 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsinfecdis.8b00276
- Primary Citation Related Structures:
6CN8 - PubMed Abstract:
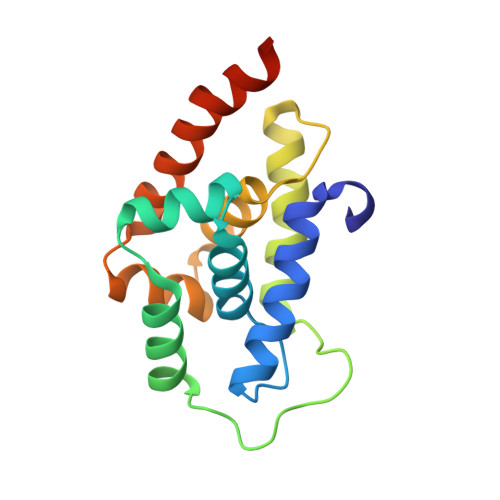
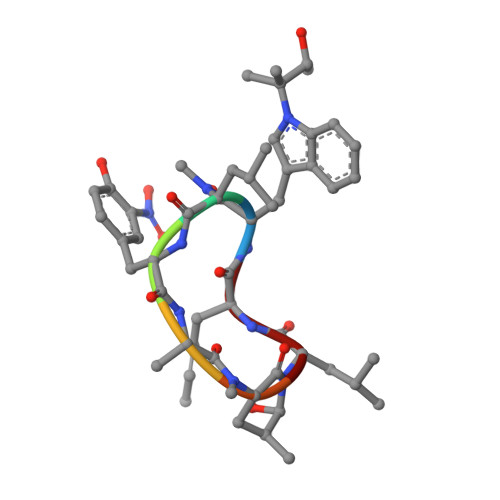
Addressing the urgent need to develop novel drugs against drug-resistant Mycobacterium tuberculosis ( M. tb) strains, ecumicin (ECU) and rufomycin I (RUFI) are being explored as promising new leads targeting cellular proteostasis via the caseinolytic protein ClpC1. Details of the binding topology and chemical mode of (inter)action of these cyclopeptides help drive further development of novel potency-optimized entities as tuberculosis drugs. ClpC1 M. tb protein constructs with mutations driving resistance to ECU and RUFI show reduced binding affinity by surface plasmon resonance (SPR). Despite certain structural similarities, ECU and RUFI resistant mutation sites did not overlap in their SPR binding patterns. SPR competition experiments show ECU prevents RUFI binding, whereas RUFI partially inhibits ECU binding. The X-ray structure of the ClpC1-NTD-RUFI complex reveals distinct differences compared to the previously reported ClpC1-NTD-cyclomarin A structure. Surprisingly, the complex structure revealed that the epoxide moiety of RUFI opened and covalently bound to ClpC1-NTD via the sulfur atom of Met1. Furthermore, RUFI analogues indicate that the epoxy group of RUFI is critical for binding and bactericidal activity. The outcomes demonstrate the significance of ClpC1 as a novel target and the importance of SAR analysis of identified macrocyclic peptides for drug discovery.
- Institute for Tuberculosis Research, College of Pharmacy , University of Illinois at Chicago , 833 S. Wood Street , Chicago , Illinois 60612 , United States.
Organizational Affiliation: