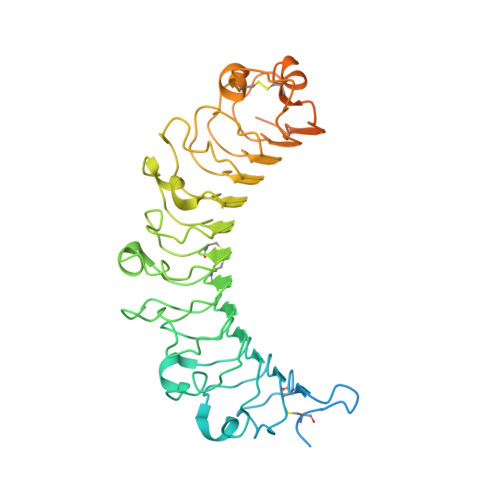
Molecular basis for governing the morphology of type-I collagen fibrils by Osteomodulin.
Tashima, T., Nagatoishi, S., Caaveiro, J.M.M., Nakakido, M., Sagara, H., Kusano-Arai, O., Iwanari, H., Mimuro, H., Hamakubo, T., Ohnuma, S.I., Tsumoto, K.(2018) Commun Biol 1: 33-33
- PubMed: 30271919 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-018-0038-2
- Primary Citation Related Structures:
5YQ5 - PubMed Abstract:
Small leucine-rich repeat proteoglycan (SLRP) proteins have an important role in the organization of the extracellular matrix, especially in the formation of collagen fibrils. However, the mechanism governing the shape of collagen fibrils is poorly understood. Here, we report that the protein Osteomodulin (OMD) of the SLRP family is a monomeric protein in solution that interacts with type-I collagen. This interaction is dominated by weak electrostatic forces employing negatively charged residues of OMD, in particular Glu284 and Glu303, and controlled by entropic factors. The protein OMD establishes a fast-binding equilibrium with collagen, where OMD may engage not only with individual collagen molecules, but also with the growing fibrils. This weak electrostatic interaction is carefully balanced so it modulates the shape of the fibrils without compromising their viability.
- Department of Chemistry & Biotechnology, School of Engineering, The University of Tokyo, Tokyo, 108-8639, Japan.
Organizational Affiliation: