Combined CRISPRi/a-Based Chemical Genetic Screens Reveal that Rigosertib Is a Microtubule-Destabilizing Agent.
Jost, M., Chen, Y., Gilbert, L.A., Horlbeck, M.A., Krenning, L., Menchon, G., Rai, A., Cho, M.Y., Stern, J.J., Prota, A.E., Kampmann, M., Akhmanova, A., Steinmetz, M.O., Tanenbaum, M.E., Weissman, J.S.(2017) Mol Cell 68: 210-223.e6
- PubMed: 28985505 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2017.09.012
- Primary Citation Related Structures:
5OV7 - PubMed Abstract:
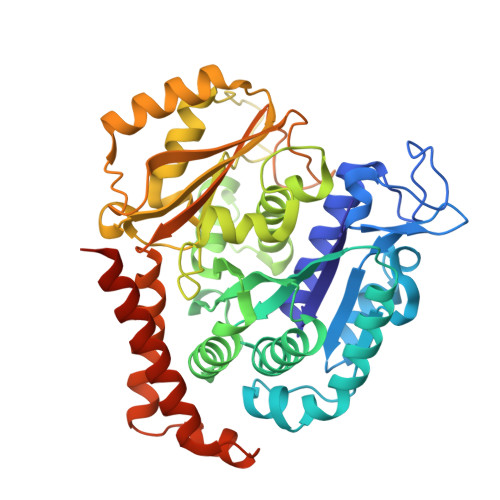
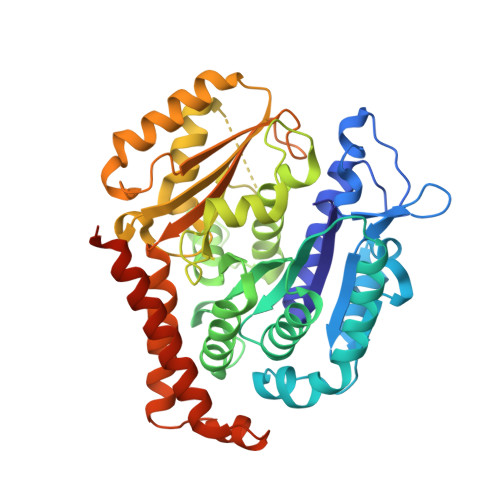
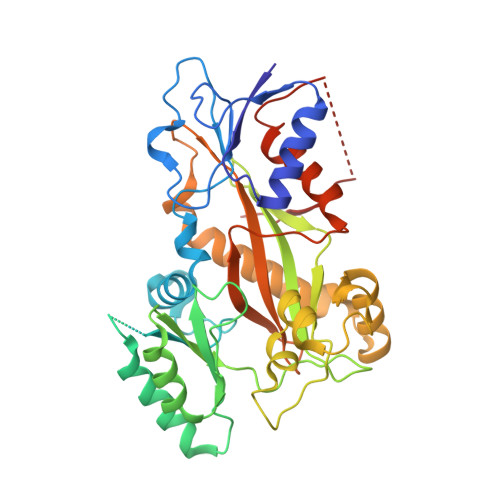
Chemical libraries paired with phenotypic screens can now readily identify compounds with therapeutic potential. A central limitation to exploiting these compounds, however, has been in identifying their relevant cellular targets. Here, we present a two-tiered CRISPR-mediated chemical-genetic strategy for target identification: combined genome-wide knockdown and overexpression screening as well as focused, comparative chemical-genetic profiling. Application of these strategies to rigosertib, a drug in phase 3 clinical trials for high-risk myelodysplastic syndrome whose molecular target had remained controversial, pointed singularly to microtubules as rigosertib's target. We showed that rigosertib indeed directly binds to and destabilizes microtubules using cell biological, in vitro, and structural approaches. Finally, expression of tubulin with a structure-guided mutation in the rigosertib-binding pocket conferred resistance to rigosertib, establishing that rigosertib kills cancer cells by destabilizing microtubules. These results demonstrate the power of our chemical-genetic screening strategies for pinpointing the physiologically relevant targets of chemical agents.
- Department of Cellular and Molecular Pharmacology, University of California, San Francisco, San Francisco, CA 94158, USA; Howard Hughes Medical Institute, University of California, San Francisco, San Francisco, CA 94158, USA; Center for RNA Systems Biology, University of California, San Francisco, San Francisco, CA 94158, USA; Department of Microbiology and Immunology, University of California, San Francisco, San Francisco, CA 94158, USA.
Organizational Affiliation: