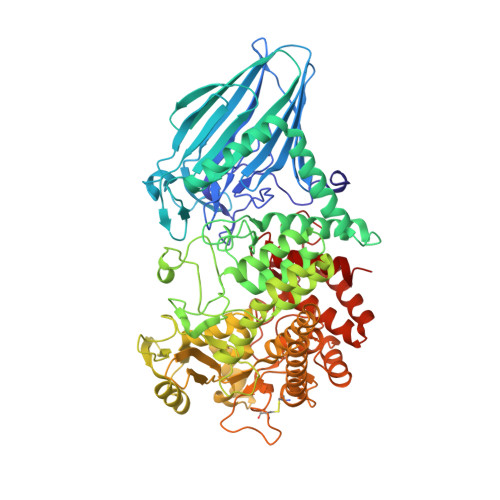
Crystal structure of the enzyme-product complex reveals sugar ring distortion during catalysis by family 63 inverting alpha-glycosidase.
Miyazaki, T., Nishikawa, A., Tonozuka, T.(2016) J Struct Biol
- PubMed: 27688023 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2016.09.015
- Primary Citation Related Structures:
5CA3, 5GW7 - PubMed Abstract:
Glycoside hydrolases are divided into two groups, known as inverting and retaining enzymes, based on their hydrolytic mechanisms. Glycoside hydrolase family 63 (GH63) is composed of inverting α-glycosidases, which act mainly on α-glucosides. We previously found that Escherichia coli GH63 enzyme, YgjK, can hydrolyze 2-O-α-d-glucosyl-d-galactose. Two constructed glycosynthase mutants, D324N and E727A, which catalyze the transfer of a β-glucosyl fluoride donor to galactose, lactose, and melibiose. Here, we determined the crystal structures of D324N and E727A soaked with a mixture of glucose and lactose at 1.8- and 2.1-Å resolutions, respectively. Because glucose and lactose molecules are found at the active sites in both structures, it is possible that these structures mimic the enzyme-product complex of YgjK. A glucose molecule found at subsite -1 in both structures adopts an unusual 1 S 3 skew-boat conformation. Comparison between these structures and the previously determined enzyme-substrate complex structure reveals that the glucose pyranose ring might be distorted immediately after nucleophilic attack by a water molecule. These structures represent the first enzyme-product complex for the GH63 family, as well as the structurally-related glycosidases, and it may provide insight into the catalytic mechanism of these enzymes.
- Research Institute of Green Science and Technology, Shizuoka University, 836 Ohya, Suruga-ku, Shizuoka 422-8529 Japan; Department of Applied Biological Science, Tokyo University of Agriculture and Technology, 3-5-8 Saiwai-cho, Fuchu, Tokyo 183-8509, Japan.
Organizational Affiliation: