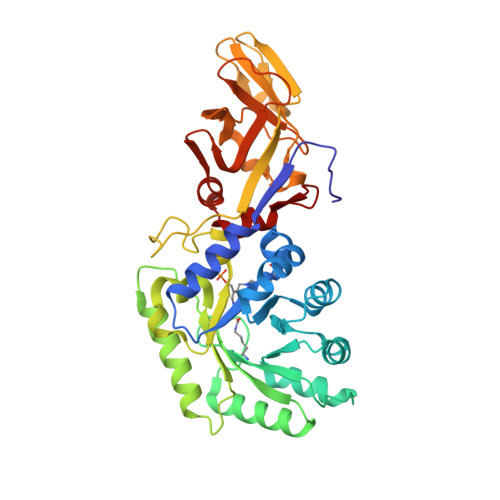
The crystal structure of alanine racemase from Streptococcus pneumoniae, a target for structure-based drug design.
Im, H., Sharpe, M.L., Strych, U., Davlieva, M., Krause, K.L.(2011) BMC Microbiol 11: 116-116
- PubMed: 21612658 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1471-2180-11-116
- Primary Citation Related Structures:
3S46 - PubMed Abstract:
Streptococcus pneumoniae is a globally important pathogen. The Gram-positive diplococcus is a leading cause of pneumonia, otitis media, bacteremia, and meningitis, and antibiotic resistant strains have become increasingly common over recent years. Alanine racemase is a ubiquitous enzyme among bacteria and provides the essential cell wall precursor, D-alanine. Since it is absent in humans, this enzyme is an attractive target for the development of drugs against S. pneumoniae and other bacterial pathogens. Here we report the crystal structure of alanine racemase from S. pneumoniae (AlrSP). Crystals diffracted to a resolution of 2.0 Å and belong to the space group P3121 with the unit cell parameters a = b = 119.97 Å, c = 118.10 Å, α = β = 90° and γ = 120°. Structural comparisons show that AlrSP shares both an overall fold and key active site residues with other bacterial alanine racemases. The active site cavity is similar to other Gram positive alanine racemases, featuring a restricted but conserved entryway. We have solved the structure of AlrSP, an essential step towards the development of an accurate pharmacophore model of the enzyme, and an important contribution towards our on-going alanine racemase structure-based drug design project. We have identified three regions on the enzyme that could be targeted for inhibitor design, the active site, the dimer interface, and the active site entryway.
- Research Institute of Pharmaceutical Sciences, College of Pharmacy, Seoul National University, Seoul, Korea.
Organizational Affiliation: