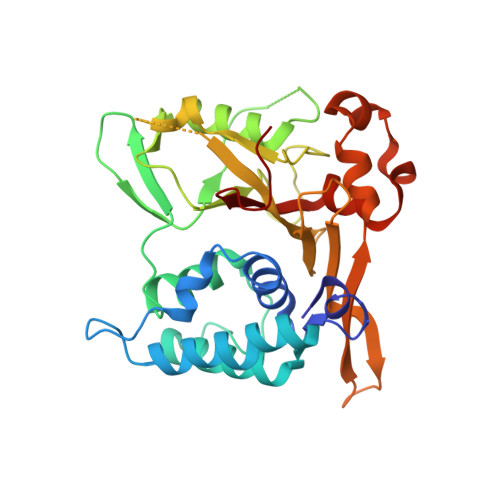
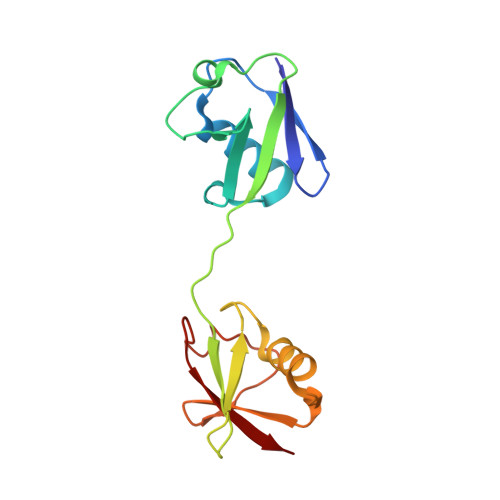
Structures of CYLD USP with Met1- or Lys63-linked diubiquitin reveal mechanisms for dual specificity.
Sato, Y., Goto, E., Shibata, Y., Kubota, Y., Yamagata, A., Goto-Ito, S., Kubota, K., Inoue, J., Takekawa, M., Tokunaga, F., Fukai, S.(2015) Nat Struct Mol Biol 22: 222-229
- PubMed: 25686088 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2970
- Primary Citation Related Structures:
3WXE, 3WXF, 3WXG - PubMed Abstract:
The tumor suppressor CYLD belongs to a ubiquitin (Ub)-specific protease (USP) family and specifically cleaves Met1- and Lys63-linked polyubiquitin chains to suppress inflammatory signaling pathways. Here, we report crystal structures representing the catalytic states of zebrafish CYLD for Met1- and Lys63-linked Ub chains and two distinct precatalytic states for Met1-linked chains. In both catalytic states, the distal Ub is bound to CYLD in a similar manner, and the scissile bond is located close to the catalytic residue, whereas the proximal Ub is bound in a manner specific to Met1- or Lys63-linked chains. Further structure-based mutagenesis experiments support the mechanism by which CYLD specifically cleaves both Met1- and Lys63-linked chains and provide insight into tumor-associated mutations of CYLD. This study provides new structural insight into the mechanisms by which USP family deubiquitinating enzymes recognize and cleave Ub chains with specific linkage types.
- 1] Life Science Division, Synchrotron Radiation Research Organization, University of Tokyo, Tokyo, Japan. [2] Center for Structural Biology of Challenging Proteins, Institute of Molecular and Cellular Biosciences, University of Tokyo, Tokyo, Japan. [3] Department of Medical Genome Sciences, Graduate School of Frontier Sciences, University of Tokyo, Chiba, Japan.
Organizational Affiliation: