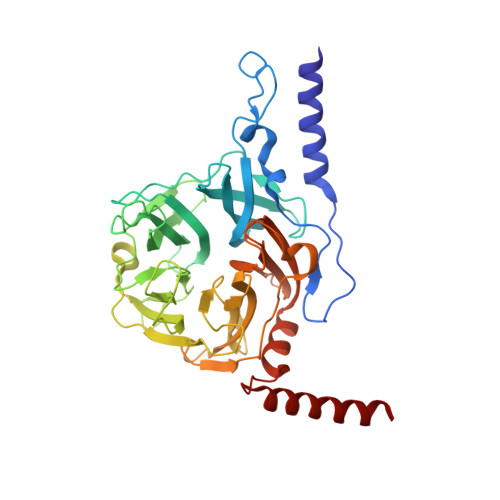
Structure of Novel Enzyme in Mannan Biodegradation Process 4-O-beta-d-Mannosyl-d-Glucose Phosphorylase MGP
Nakae, S., Ito, S., Higa, M., Senoura, T., Wasaki, J., Hijikata, A., Shionyu, M., Ito, S., Shirai, T.(2013) J Mol Biology 425: 4468-4478
- PubMed: 23954514 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2013.08.002
- Primary Citation Related Structures:
3WAS, 3WAT, 3WAU, 4KMI - PubMed Abstract:
The crystal structure of a novel component of the mannan biodegradation system, 4-O-β-D-mannosyl-D-glucose phosphorylase (MGP), was determined to a 1.68-Å resolution. The structure of the enzyme revealed a unique homohexameric structure, which was formed by using two helices attached to the N-terminus and C-terminus as a tab for sticking between subunits. The structures of MGP complexes with genuine substrates, 4-O-β-D-mannosyl-D-glucose and phosphate, and the product D-mannose-1-phosphate were also determined. The complex structures revealed that the invariant residue Asp131, which is supposed to be the general acid/base, did not exist close to the glycosidic Glc-O4 atom, which should be protonated in the catalytic reaction. Also, no solvent molecule that might mediate a proton transfer from Asp131 was observed in the substrate complex structure, suggesting that the catalytic mechanism of MGP is different from those of known disaccharide phosphorylases.
- Department of Bioscience, Nagahama Institute of Bio-science and Technology, and Japan Science and Technology Agency Institute for Bioinformatics Research and Development, 1266 Tamura, Nagahama 526-0829, Japan.
Organizational Affiliation: