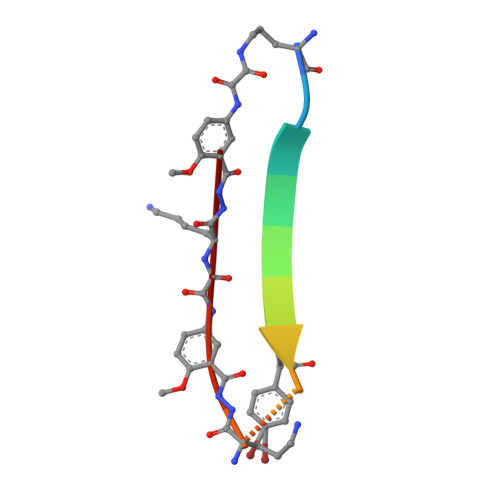
X-ray crystallographic structure of an artificial beta-sheet dimer.
Khakshoor, O., Lin, A.J., Korman, T.P., Sawaya, M.R., Tsai, S.C., Eisenberg, D., Nowick, J.S.(2010) J Am Chem Soc 132: 11622-11628
- PubMed: 20669960 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja103438w
- Primary Citation Related Structures:
3NI3 - PubMed Abstract:
This paper describes the X-ray crystallographic structure of a designed cyclic beta-sheet peptide that forms a well-defined hydrogen-bonded dimer that mimics beta-sheet dimers formed by proteins. The 54-membered ring macrocyclic peptide (1a) contains molecular template and turn units that induce beta-sheet structure in a heptapeptide strand that forms the dimerization interface. The X-ray crystallographic structure reveals the structures of the two "Hao" amino acids that help template the beta-sheet structure and the two delta-linked ornithine turn units that link the Hao-containing template to the heptapeptide beta-strand. The Hao amino acids adopt a conformation that resembles a tripeptide in a beta-strand conformation, with one edge of the Hao unit presenting an alternating array of hydrogen-bond donor and acceptor groups in the same pattern as that of a tripeptide beta-strand. The delta-linked ornithines adopt a conformation that resembles a hydrogen-bonded beta-turn, in which the ornithine takes the place of the i+1 and i+2 residues. The dimers formed by macrocyclic beta-sheet 1a resemble the dimers of many proteins, such as defensin HNP-3, the lambda-Cro repressor, interleukin 8, and the ribonuclease H domain of HIV-1 reverse transcriptase. The dimers of 1a self-assemble in the solid state into a barrel-shaped trimer of dimers in which the three dimers are arranged in a triangular fashion. Molecular modeling in which one of the three dimers is removed and the remaining two dimers are aligned face-to-face provides a model of the dimers of dimers of closely related macrocyclic beta-sheet peptides that were observed in solution.
- Department of Chemistry, University of California, Irvine, California 92697-2025, USA.
Organizational Affiliation: