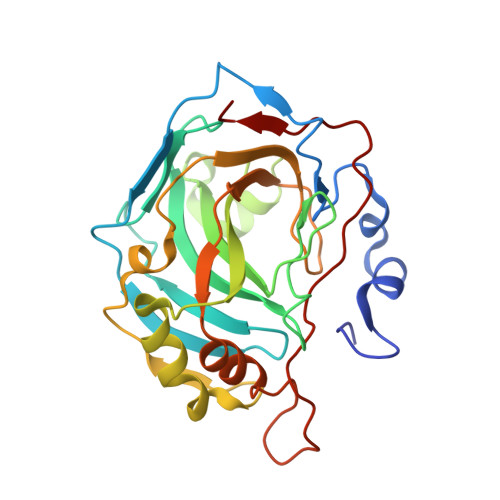
Entrapment of carbon dioxide in the active site of carbonic anhydrase II
Domsic, J.F., Avvaru, B.S., Kim, C.U., Gruner, S.M., Agbandje-McKenna, M., Silverman, D.N., McKenna, R.(2008) J Biological Chem 283: 30766-30771
- PubMed: 18768466 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M805353200
- Primary Citation Related Structures:
3D92, 3D93 - PubMed Abstract:
The visualization at near atomic resolution of transient substrates in the active site of enzymes is fundamental to fully understanding their mechanism of action. Here we show the application of using CO(2)-pressurized, cryo-cooled crystals to capture the first step of CO(2) hydration catalyzed by the zinc-metalloenzyme human carbonic anhydrase II, the binding of substrate CO(2), for both the holo and the apo (without zinc) enzyme to 1.1A resolution. Until now, the feasibility of such a study was thought to be technically too challenging because of the low solubility of CO(2) and the fast turnover to bicarbonate by the enzyme (Liang, J. Y., and Lipscomb, W. N. (1990) Proc. Natl. Acad. Sci. U. S. A. 87, 3675-3679). These structures provide insight into the long hypothesized binding of CO(2) in a hydrophobic pocket at the active site and demonstrate that the zinc does not play a critical role in the binding or orientation of CO(2). This method may also have a much broader implication for the study of other enzymes for which CO(2) is a substrate or product and for the capturing of transient substrates and revealing hydrophobic pockets in proteins.
- Department of Biochemistry and Molecular Biology, University of Florida, Gainesville, Florida 32610, USA.
Organizational Affiliation: