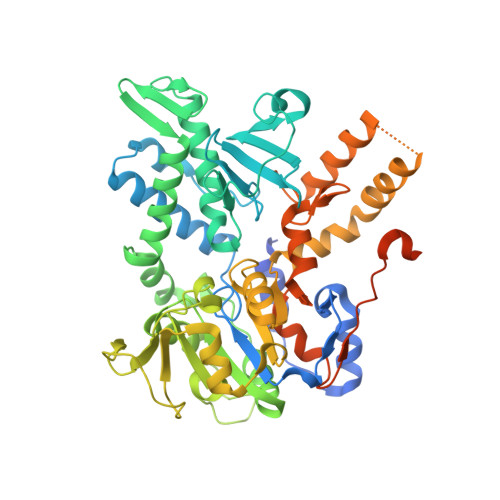
Structural Basis for the Allosteric Regulation and Substrate Recognition of Human Cytosolic 5'-Nucleotidase II
Wallden, K., Nordlund, P.(2011) J Mol Biology 408: 684
- PubMed: 21396942
- DOI: https://doi.org/10.1016/j.jmb.2011.02.059
- Primary Citation Related Structures:
2XCV, 2XCW, 2XCX, 2XJB, 2XJC, 2XJD, 2XJE, 2XJF - PubMed Abstract:
Cytosolic 5'-nucleotidase II (cN-II) catalyzes the dephosphorylation of 6-hydroxypurine nucleoside 5'-monophosphates and participates in the regulation of purine nucleotide pools within the cell. It interferes with the phosphorylation-dependent activation of nucleoside analogues used in the treatment of cancer and viral diseases. It is allosterically activated by a number of phosphate-containing cellular metabolites such as ATP, diadenosine polyphosphates, and 2,3-bisphosphoglycerate, which couple its activity with the metabolic state of the cell. We present seven high-resolution structures of human cN-II, including a ligand-free form and complexes with various substrates and effectors. These structures reveal the structural basis for the allosteric activation of cN-II, uncovering a mechanism where an effector-induced disorder-to-order transition generates rearrangements within the catalytic site and the subsequent coordination of the catalytically essential magnesium. Central to the activation is the large transition of the catalytically essential Asp356. This study also provides the structural basis for the substrate specificity of cN-II, where Arg202, Asp206, and Phe157 seem to be important residues for purine/pyrimidine selectivity. These structures provide a comprehensive structural basis for the design of cN-II inhibitors. They also contribute to the understanding of how the nucleotide salvage pathway is regulated at a molecular level.
- Department of Medical Biochemistry and Biophysics, Karolinska Institutet, 17177 Stockholm, Sweden.
Organizational Affiliation: