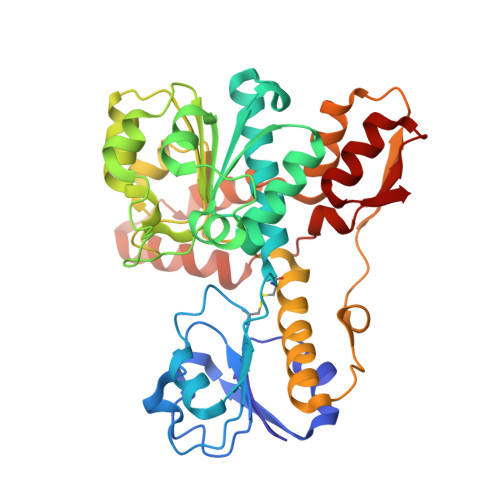
Crystal Structure of d-Erythronate-4-phosphate Dehydrogenase Complexed with NAD
Ha, J.Y., Lee, J.H., Kim, K.H., Kim, D.J., Lee, H.H., Kim, H.K., Yoon, H.J., Suh, S.W.(2007) J Mol Biology 366: 1294-1304
- PubMed: 17217963
- DOI: https://doi.org/10.1016/j.jmb.2006.12.038
- Primary Citation of Related Structures:
2O4C - PubMed Abstract:
Pyridoxal-5'-phosphate (the active form of vitamin B6) is an essential cofactor in many enzymatic reactions. While animals lack any of the pathways for de novo synthesis and salvage of vitamin B6, it is synthesized by two distinct biosynthetic routes in bacteria, fungi, parasites, and plants. One of them is the PdxA/PdxJ pathway found in the gamma subdivision of proteobacteria. It depends on the pdxB gene, which encodes erythronate-4-phosphate dehydrogenase (PdxB), a member of the d-isomer specific 2-hydroxyacid dehydrogenase superfamily. Although three-dimensional structures of other functionally related dehydrogenases are available, no structure of PdxB has been reported. To provide the missing structural information and to gain insights into the catalytic mechanism, we have determined the first crystal structure of erythronate-4-phosphate dehydrogenase from Pseudomonas aeruginosa in the ligand-bound state. It is a homodimeric enzyme consisting of 380-residue subunits. Each subunit consists of three structural domains: the lid domain, the nucleotide-binding domain, and the C-terminal dimerization domain. The latter domain has a unique fold and is largely responsible for dimerization. Interestingly, two subunits of the dimeric enzyme are bound with different combinations of ligands in the crystal and they display significantly different conformations. Subunit A is bound with NAD and a phosphate ion, while subunit B, with a more open active site cleft, is bound with NAD and l(+)-tartrate. Our structural data allow a detailed understanding of cofactor and substrate recognition, thus providing substantial insights into PdxB catalysis.
- Department of Chemistry, College of Natural Sciences, Seoul National University, Seoul 151-742, Korea.
Organizational Affiliation: