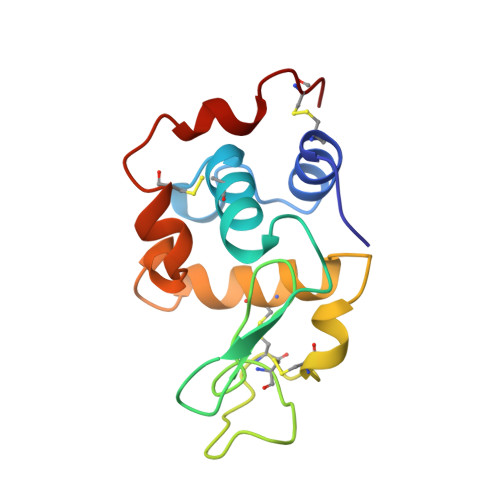
Structures of monoclinic lysozyme iodide at 1.6 A and of triclinic lysozyme nitrate at 1.1 A.
Steinrauf, L.K.(1998) Acta Crystallogr D Biol Crystallogr 54: 767-780
- PubMed: 9757091 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444997016922
- Primary Citation Related Structures:
1LKR, 1LKS - PubMed Abstract:
Hen egg-white lysozyme is one of the most thoroughly studied of enzymes and has been the subject of study by many methods, including X-ray crystallography. The present work extends the X-ray crystallography to higher resolution, includes the positions of the anions, and examines the contacts of the neighbors in greater detail. Data were collected at room temperature on a Rigaku R-axis area detector with rotating-anode X-ray generator to 1.6 A resolution for monoclinic lysozyme iodide at pH 4.0, to 1.8 A for monoclinic lysozyme iodide at pH 8.0, and to 1.1 A resolution for triclinic lysozyme nitrate at pH 4.5. The structures have been refined by SHELX93 with the expected number of anion sites being accounted for. Two regions of the protein have been found to be variable: residues 65-75 and 99-104. Except for 65-75 and 99-104, lysozyme is a very stable molecule with the crystal forms being held together by the electrostatic contacts of the anions and by layers of water molecules. The anion positions can be described as paired half sites, each half being contributed by a different lysozyme molecule. The many different crystal forms of lysozyme may be due to different combinations of the many such half sites on the surface. A hypothesis is presented for lysozyme in the different crystal forms and which may be extended to behavior in solution. Suggestions for future crystallographic research are proposed, involving anions of different shape and charge.
- Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis, IN 46202-5122, USA. lsteinra@mail.nsysu.edu.tw
Organizational Affiliation: