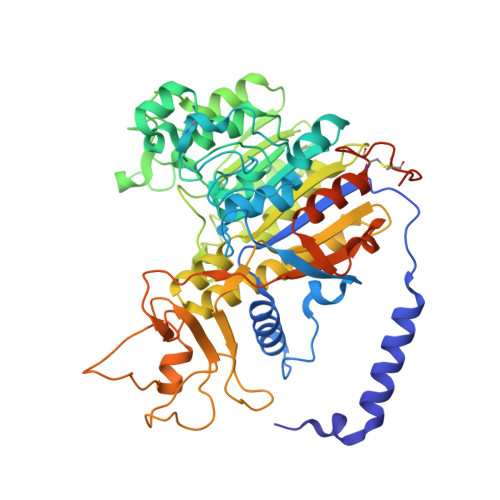
Crystal structure of alkaline phosphatase from human placenta at 1.8 A resolution. Implication for a substrate specificity.
Le Du, M.H., Stigbrand, T., Taussig, M.J., Menez, A., Stura, E.A.(2001) J Biological Chem 276: 9158-9165
- PubMed: 11124260 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M009250200
- Primary Citation Related Structures:
1EW2 - PubMed Abstract:
Human placental alkaline phosphatase (PLAP) is one of three tissue-specific human APs extensively studied because of its ectopic expression in tumors. The crystal structure, determined at 1.8-A resolution, reveals that during evolution, only the overall features of the enzyme have been conserved with respect to Escherichia coli. The surface is deeply mutated with 8% residues in common, and in the active site, only residues strictly necessary to perform the catalysis have been preserved. Additional structural elements aid an understanding of the allosteric property that is specific for the mammalian enzyme (Hoylaerts, M. F., Manes, T., and Millán, J. L. (1997) J. Biol. Chem. 272, 22781-22787). Allostery is probably favored by the quality of the dimer interface, by a long N-terminal alpha-helix from one monomer that embraces the other one, and similarly by the exchange of a residue from one monomer in the active site of the other. In the neighborhood of the catalytic serine, the orientation of Glu-429, a residue unique to PLAP, and the presence of a hydrophobic pocket close to the phosphate product, account for the specific uncompetitive inhibition of PLAP by l-amino acids, consistent with the acquisition of substrate specificity. The location of the active site at the bottom of a large valley flanked by an interfacial crown-shaped domain and a domain containing an extra metal ion on the other side suggest that the substrate of PLAP could be a specific phosphorylated protein.
- Département d'Ingénierie et d'Etudes des Protéines (DIEP), Commissariat à l'Energie Atomique, C. E. Saclay, 91191 Gif-sur-Yvette Cedex, France. mhledu@cea.fr
Organizational Affiliation: