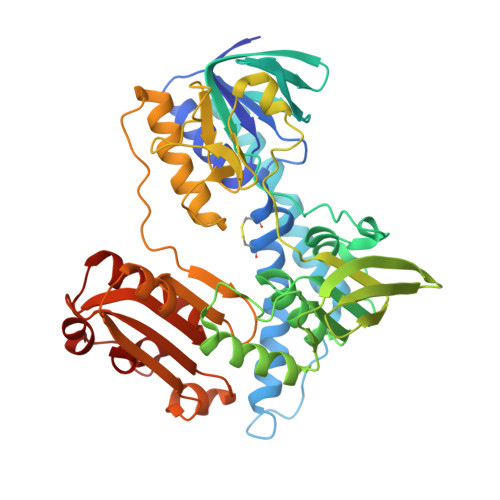
Biochemical and structural basis of mercuric reductase, GbsMerA, from Gelidibacter salicanalis PAMC21136.
Pardhe, B.D., Lee, M.J., Lee, J.H., Do, H., Oh, T.J.(2023) Sci Rep 13: 17854-17854
- PubMed: 37857791 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-023-44968-w
- Primary Citation Related Structures:
8K40, 8K41 - PubMed Abstract:
Heavy metals, including mercury, are non-biodegradable and highly toxic to microorganisms even at low concentrations. Understanding the mechanisms underlying the environmental adaptability of microorganisms with Hg resistance holds promise for their use in Hg bioremediation. We characterized GbsMerA, a mercury reductase belonging to the mercury-resistant operon of Gelidibacter salicanalis PAMC21136, and found its maximum activity of 474.7 µmol/min/mg in reducing Hg +2 . In the presence of Ag and Mn, the enzyme exhibited moderate activity as 236.5 µmol/min/mg and 69 µmol/min/mg, respectively. GbsMerA exhibited optimal activity at pH 7.0 and a temperature of 60 °C. Moreover, the crystal structure of GbsMerA and structural comparison with homologues indicated that GbsMerA contains residues, Tyr437´ and Asp47, which may be responsible for metal transfer at the si-face by providing a hydroxyl group (-OH) to abstract a proton from the thiol group of cysteine. The complex structure with NADPH indicated that Y174 in the re-face can change its side chain direction upon NADPH binding, indicating that Y174 may have a role as a gate for NADPH binding. Moreover, the heterologous host expressing GbsMerA (pGbsMerA) is more resistant to Hg toxicity when compared to the host lacking GbsMerA. Overall, this study provides a background for understanding the catalytic mechanism and Hg detoxification by GbsMerA and suggests the application of genetically engineered E. coli strains for environmental Hg removal.
- Department of Life Science and Biochemical Engineering, Graduate School, SunMoon University, Asan, 31460, Republic of Korea.
Organizational Affiliation: