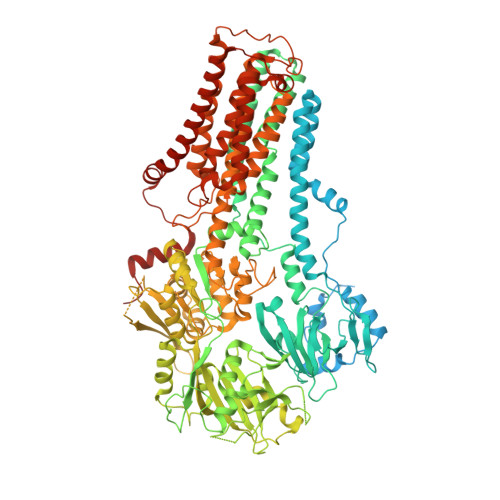
Cryo-EM reveals mechanistic insights into lipid-facilitated polyamine export by human ATP13A2.
Tomita, A., Daiho, T., Kusakizako, T., Yamashita, K., Ogasawara, S., Murata, T., Nishizawa, T., Nureki, O.(2021) Mol Cell 81: 4799-4809.e5
- PubMed: 34798056 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2021.11.001
- Primary Citation Related Structures:
7VPI, 7VPJ, 7VPK, 7VPL - PubMed Abstract:
The cytoplasmic polyamine maintains cellular homeostasis by chelating toxic metal cations, regulating transcriptional activity, and protecting DNA. ATP13A2 was identified as a lysosomal polyamine exporter responsible for polyamine release into the cytosol, and its dysfunction is associated with Alzheimer's disease and other neural degradation diseases. ATP13A2 belongs to the P5 subfamily of the P-type ATPase family, but its mechanisms remain unknown. Here, we report the cryoelectron microscopy (cryo-EM) structures of human ATP13A2 under four different conditions, revealing the structural coupling between the polyamine binding and the dephosphorylation. Polyamine is bound at the luminal tunnel and recognized through numerous electrostatic and π-cation interactions, explaining its broad specificity. The unique N-terminal domain is anchored to the lipid membrane to stabilize the E2P conformation, thereby accelerating the E1P-to-E2P transition. These findings reveal the distinct mechanism of P5B ATPases, thereby paving the way for neuroprotective therapy by activating ATP13A2.
- Department of Biological Sciences, Graduate School of Science, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: