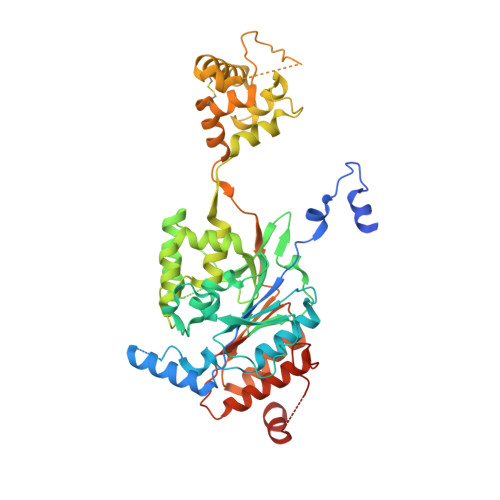
Structures of the mannose-6-phosphate pathway enzyme, GlcNAc-1-phosphotransferase.
Gorelik, A., Illes, K., Bui, K.H., Nagar, B.(2022) Proc Natl Acad Sci U S A 119: e2203518119-e2203518119
- PubMed: 35939698
- DOI: https://doi.org/10.1073/pnas.2203518119
- Primary Citation Related Structures:
7S69, 7S6N, 7SJ2 - PubMed Abstract:
The mannose-6-phosphate (M6P) pathway is responsible for the transport of hydrolytic enzymes to lysosomes. N-acetylglucosamine-1-phosphotransferase (GNPT) catalyzes the first step of tagging these hydrolases with M6P, which when recognized by receptors in the Golgi diverts them to lysosomes. Genetic defects in the GNPT subunits, GNPTAB and GNPTG, cause the lysosomal storage diseases mucolipidosis types II and III. To better understand its function, we determined partial three-dimensional structures of the GNPT complex. The catalytic domain contains a deep cavity for binding of uridine diphosphate- N -acetylglucosamine, and the surrounding residues point to a one-step transfer mechanism. An isolated structure of the gamma subunit of GNPT reveals that it can bind to mannose-containing glycans in different configurations, suggesting that it may play a role in directing glycans into the active site. These findings may facilitate the development of therapies for lysosomal storage diseases.
- Department of Biochemistry, McGill University, Montreal, QC H3G 0B1, Canada.
Organizational Affiliation: