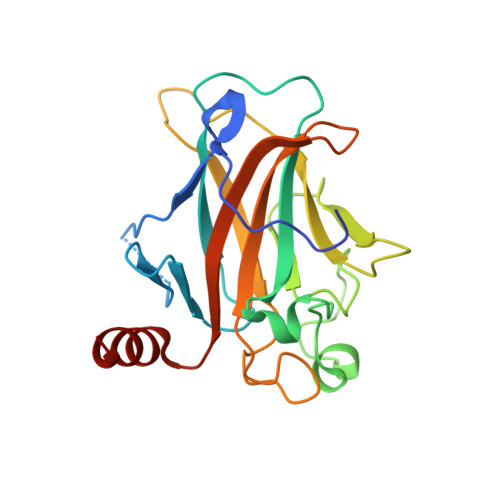
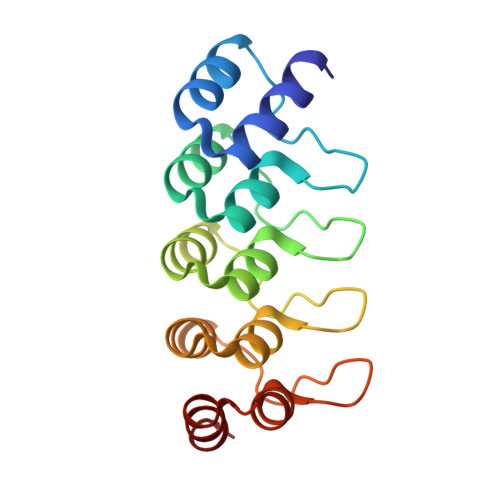
Designed Ankyrin Repeat Proteins as a tool box for analyzing p63.
Strubel, A., Munick, P., Chaikuad, A., Dreier, B., Schaefer, J., Gebel, J., Osterburg, C., Tuppi, M., Schafer, B., Knapp, S., Pluckthun, A., Dotsch, V.(2022) Cell Death Differ 29: 2445-2458
- PubMed: 35717504 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41418-022-01030-y
- Primary Citation Related Structures:
7Z71, 7Z72, 7Z73, 7Z7E - PubMed Abstract:
The function of the p53 transcription factor family is dependent on several folded domains. In addition to a DNA-binding domain, members of this family contain an oligomerization domain. p63 and p73 also contain a C-terminal Sterile α-motif domain. Inhibition of most transcription factors is difficult as most of them lack deep pockets that can be targeted by small organic molecules. Genetic knock-out procedures are powerful in identifying the overall function of a protein, but they do not easily allow one to investigate roles of individual domains. Here we describe the characterization of Designed Ankyrin Repeat Proteins (DARPins) that were selected as tight binders against all folded domains of p63. We determine binding affinities as well as specificities within the p53 protein family and show that DARPins can be used as intracellular inhibitors for the modulation of transcriptional activity. By selectively inhibiting DNA binding of the ΔNp63α isoform that competes with p53 for the same promoter sites, we show that p53 can be reactivated. We further show that inhibiting the DNA binding activity stabilizes p63, thus providing evidence for a transcriptionally regulated negative feedback loop. Furthermore, the ability of DARPins to bind to the DNA-binding domain and the Sterile α-motif domain within the dimeric-only and DNA-binding incompetent conformation of TAp63α suggests a high structural plasticity within this special conformation. In addition, the developed DARPins can also be used to specifically detect p63 in cell culture and in primary tissue and thus constitute a very versatile research tool for studying the function of p63.
- Institute of Biophysical Chemistry and Center for Biomolecular Magnetic Resonance, Goethe University, 60438, Frankfurt, Germany.
Organizational Affiliation: