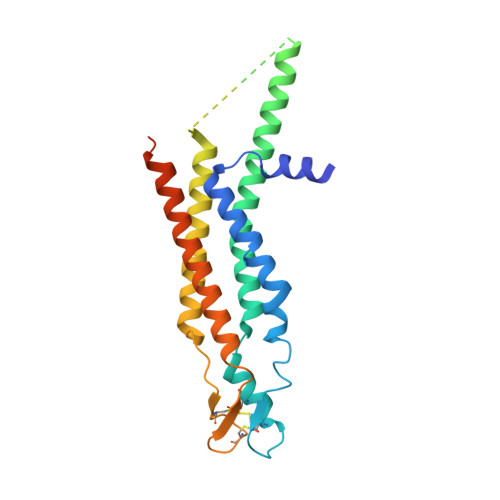
Conformational changes in the human Cx43/GJA1 gap junction channel visualized using cryo-EM.
Lee, H.J., Cha, H.J., Jeong, H., Lee, S.N., Lee, C.W., Kim, M., Yoo, J., Woo, J.S.(2023) Nat Commun 14: 931-931
- PubMed: 36805660 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-36593-y
- Primary Citation Related Structures:
7F92, 7F93, 7F94, 7XQ9, 7XQB, 7XQD, 7XQF, 7XQG, 7XQH, 7XQI, 7XQJ - PubMed Abstract:
Connexin family proteins assemble into hexameric hemichannels in the cell membrane. The hemichannels dock together between two adjacent membranes to form gap junction intercellular channels (GJIChs). We report the cryo-electron microscopy structures of Cx43 GJICh, revealing the dynamic equilibrium state of various channel conformations in detergents and lipid nanodiscs. We identify three different N-terminal helix conformations of Cx43-gate-covering (GCN), pore-lining (PLN), and flexible intermediate (FIN)-that are randomly distributed in purified GJICh particles. The conformational equilibrium shifts to GCN by cholesteryl hemisuccinates and to PLN by C-terminal truncations and at varying pH. While GJIChs that mainly comprise GCN protomers are occluded by lipids, those containing conformationally heterogeneous protomers show markedly different pore sizes. We observe an α-to-π-helix transition in the first transmembrane helix, which creates a side opening to the membrane in the FIN and PLN conformations. This study provides basic structural information to understand the mechanisms of action and regulation of Cx43 GJICh.
- Department of Life Sciences, Korea University, Seoul, 02841, Korea.
Organizational Affiliation: