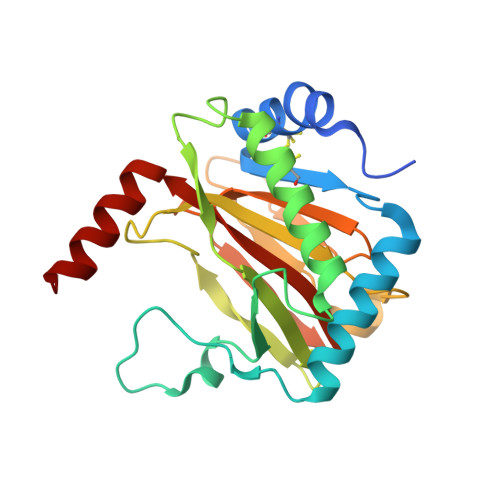
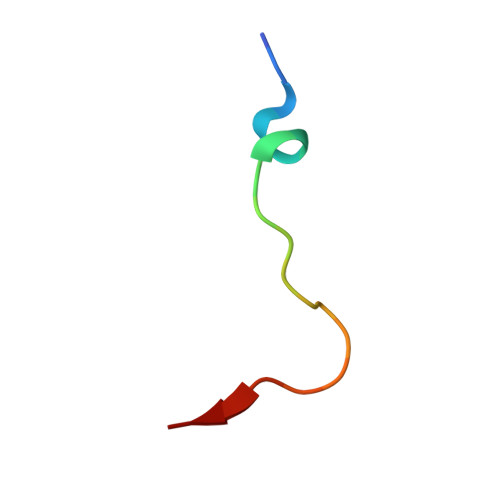
Structural basis for binding of the renal carcinoma target hypoxia-inducible factor 2 alpha to prolyl hydroxylase domain 2.
Figg Jr., W.D., Fiorini, G., Chowdhury, R., Nakashima, Y., Tumber, A., McDonough, M.A., Schofield, C.J.(2023) Proteins 91: 1510-1524
- PubMed: 37449559 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.26541
- Primary Citation Related Structures:
7Q5V, 7Q5X - PubMed Abstract:
The hypoxia-inducible factor (HIF) prolyl-hydroxylases (human PHD1-3) catalyze prolyl hydroxylation in oxygen-dependent degradation (ODD) domains of HIFα isoforms, modifications that signal for HIFα proteasomal degradation in an oxygen-dependent manner. PHD inhibitors are used for treatment of anemia in kidney disease. Increased erythropoietin (EPO) in patients with familial/idiopathic erythrocytosis and pulmonary hypertension is associated with mutations in EGLN1 (PHD2) and EPAS1 (HIF2α); a drug inhibiting HIF2α activity is used for clear cell renal cell carcinoma (ccRCC) treatment. We report crystal structures of PHD2 complexed with the C-terminal HIF2α-ODD in the presence of its 2-oxoglutarate cosubstrate or N-oxalylglycine inhibitor. Combined with the reported PHD2.HIFα-ODD structures and biochemical studies, the results inform on the different PHD.HIFα-ODD binding modes and the potential effects of clinically observed mutations in HIFα and PHD2 genes. They may help enable new therapeutic avenues, including PHD isoform-selective inhibitors and sequestration of HIF2α by the PHDs for ccRCC treatment.
- Chemistry Research Laboratory, Department of Chemistry and the Ineos Oxford, Institute for Antimicrobial Research, University of Oxford, Oxford, UK.
Organizational Affiliation: