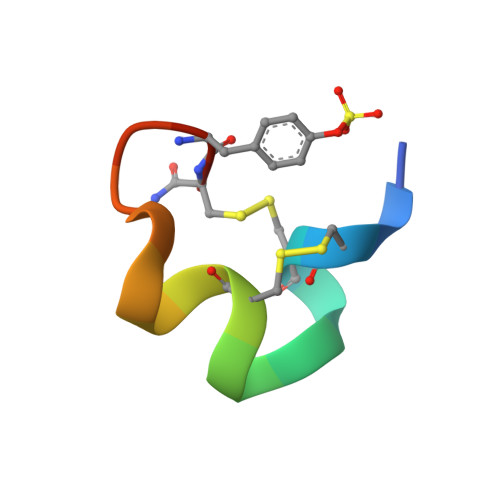
Posttranslational modifications of alpha-conotoxins: sulfotyrosine and C-terminal amidation stabilise structures and increase acetylcholine receptor binding.
Ho, T.N.T., Lee, H.S., Swaminathan, S., Goodwin, L., Rai, N., Ushay, B., Lewis, R.J., Rosengren, K.J., Conibear, A.C.(2021) RSC Med Chem 12: 1574-1584
- PubMed: 34671739 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/d1md00182e
- Primary Citation Related Structures:
7N0T, 7N1Z, 7N20, 7N21, 7N22, 7N23, 7N24, 7N25, 7N26 - PubMed Abstract:
Conotoxins are peptides found in the venoms of marine cone snails. They are typically highly structured and stable and have potent activities at nicotinic acetylcholine receptors, which make them valuable research tools and promising lead molecules for drug development. Many conotoxins are also highly modified with posttranslational modifications such as proline hydroxylation, glutamic acid gamma-carboxylation, tyrosine sulfation and C-terminal amidation, amongst others. The role of these posttranslational modifications is poorly understood, and it is unclear whether the modifications interact directly with the binding site, alter conotoxin structure, or both. Here we synthesised a set of twelve conotoxin variants bearing posttranslational modifications in the form of native sulfotyrosine and C-terminal amidation and show that these two modifications in combination increase their activity at nicotinic acetylcholine receptors and binding to soluble acetylcholine binding proteins, respectively. We then rationalise how these functional differences between variants might arise from stabilization of the three-dimensional structures and interactions with the binding sites, using high-resolution nuclear magnetic resonance data. This study demonstrates that posttranslational modifications can modulate interactions between a ligand and receptor by a combination of structural and binding alterations. A deeper mechanistic understanding of the role of posttranslational modifications in structure-activity relationships is essential for understanding receptor biology and could help to guide structure-based drug design.
- Institute for Molecular Bioscience, The University of Queensland St Lucia 4072 Brisbane Australia.
Organizational Affiliation: