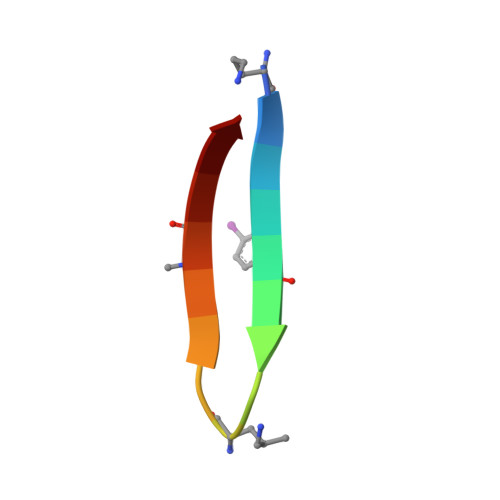
An Improved Turn Structure for Inducing beta-Hairpin Formation in Peptides.
Li, X., Sabol, A.L., Wierzbicki, M., Salveson, P.J., Nowick, J.S.(2021) Angew Chem Int Ed Engl 60: 22776-22782
- PubMed: 34258835 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.202105559
- Primary Citation Related Structures:
7LIB - PubMed Abstract:
Although β-hairpins are widespread in proteins, there is no tool to coax any small peptide to adopt a β-hairpin conformation, regardless of sequence. Here, we report that δ-linked γ(R)-methyl-ornithine ( δ MeOrn) provides an improved β-turn template for inducing a β-hairpin conformation in peptides. We developed a synthesis of protected δ MeOrn as a building block suitable for use in Fmoc-based solid-phase peptide synthesis. The synthesis begins with l-leucine and affords gram quantities of the N α -Boc-N δ -Fmoc-γ(R)-methyl-ornithine building block. X-ray crystallography confirms that the δ MeOrn turn unit adopts a folded structure in a macrocyclic β-hairpin peptide. CD and NMR spectroscopy allow comparison of the δ MeOrn turn template to the δ-linked ornithine ( δ Orn) turn template that we previously introduced and to the popular d-Pro-Gly turn template. These studies show that the folding of the δ MeOrn turn template is substantially better than that of δ Orn and is comparable to d-Pro-Gly.
- Department of Chemistry, University of California Irvine, 4126 Natural Sciences I, Irvine, CA, 92697-2025, USA.
Organizational Affiliation: