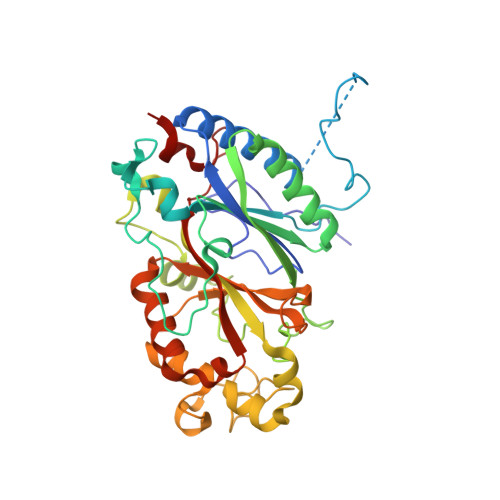
Structural insights at acidic pH of dye-decolorizing peroxidase from Bacillus subtilis.
Dhankhar, P., Dalal, V., Sharma, A.K., Kumar, P.(2022) Proteins
- PubMed: 36345957 Search on PubMed
- DOI: https://doi.org/10.1002/prot.26444
- Primary Citation Related Structures:
7E5Q - PubMed Abstract:
Dye-decolorizing peroxidases (DyPs), a type of heme-containing oxidoreductase enzymes, catalyze the peroxide-dependent oxidation of various industrial dyes as well as lignin and lignin model compounds. In our previous work, we have recently reported the crystal structures of class A-type DyP from Bacillus subtilis at pH 7.0 (BsDyP7), exposing the location of three binding sites for small substrates and high redox-potential substrates. The biochemical studies revealed the optimum acidic pH for enzyme activity. In the present study, the crystal structure of BsDyP at acidic pH (BsDyP4) reveals two-monomer units stabilized by intermolecular salt bridges and a hydrogen bond network in a homo-dimeric unit. Based on the monomeric structural comparison of BsDyP4 and BsDyP7, minor differences were observed in the loop regions, that is, LI (Ala64-Gln71), LII (Glu96-Lys108), LIII (Pro117-Leu124), and LIV (Leu295-Asp303). Despite these differences, BsDyP4 adopts similar heme architecture as well as three substrate-binding sites to BsDyP7. In BsDyP4, a shift in Asp187, heme pocket residue discloses the plausible reason for optimal acidic pH for BsDyP activity. This study provides insight into the structural changes in BsDyP at acidic pH, where BsDyP is biologically active.
- Department of Biosciences and Bioengineering, Indian Institute of Technology Roorkee, Roorkee, India.
Organizational Affiliation: