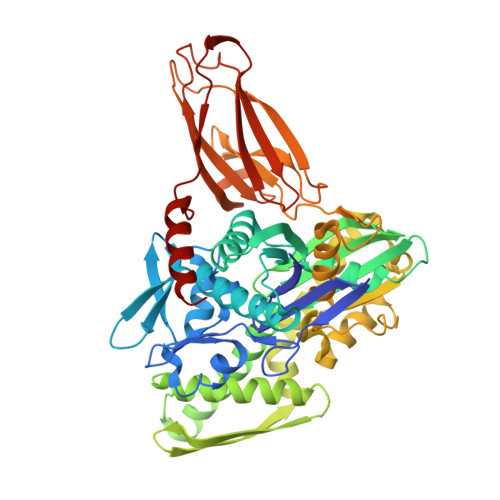
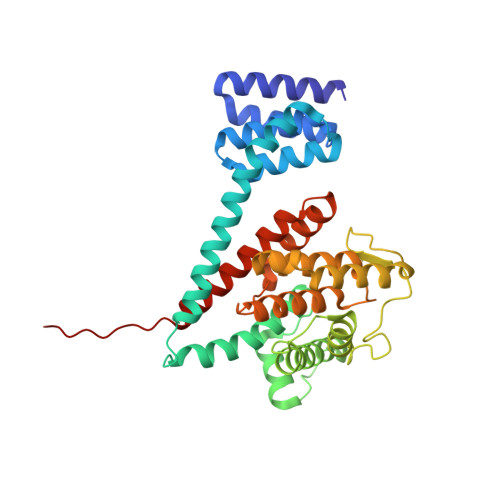
Structures of a deAMPylation complex rationalise the switch between antagonistic catalytic activities of FICD.
Perera, L.A., Preissler, S., Zaccai, N.R., Prevost, S., Devos, J.M., Haertlein, M., Ron, D.(2021) Nat Commun 12: 5004-5004
- PubMed: 34408154 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-25076-7
- Primary Citation Related Structures:
7B7Z, 7B80 - PubMed Abstract:
The endoplasmic reticulum (ER) Hsp70 chaperone BiP is regulated by AMPylation, a reversible inactivating post-translational modification. Both BiP AMPylation and deAMPylation are catalysed by a single ER-localised enzyme, FICD. Here we present crystallographic and solution structures of a deAMPylation Michaelis complex formed between mammalian AMPylated BiP and FICD. The latter, via its tetratricopeptide repeat domain, binds a surface that is specific to ATP-state Hsp70 chaperones, explaining the exquisite selectivity of FICD for BiP's ATP-bound conformation both when AMPylating and deAMPylating Thr518. The eukaryotic deAMPylation mechanism thus revealed, rationalises the role of the conserved Fic domain Glu234 as a gatekeeper residue that both inhibits AMPylation and facilitates hydrolytic deAMPylation catalysed by dimeric FICD. These findings point to a monomerisation-induced increase in Glu234 flexibility as the basis of an oligomeric state-dependent switch between FICD's antagonistic activities, despite a similar mode of engagement of its two substrates - unmodified and AMPylated BiP.
- Cambridge Institute for Medical Research, University of Cambridge, Cambridge, UK. lp397@cam.ac.uk.
Organizational Affiliation: