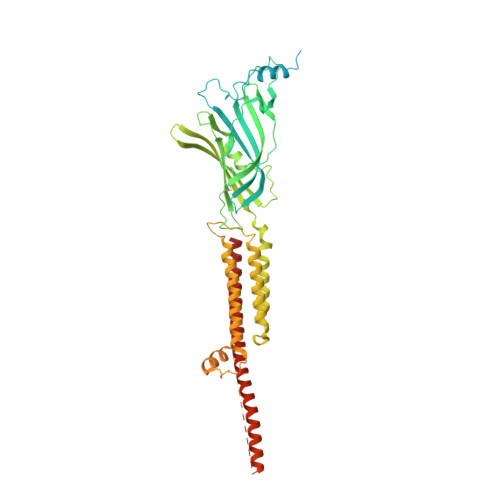
The Binding of Palonosetron and Other Antiemetic Drugs to the Serotonin 5-HT3 Receptor.
Zarkadas, E., Zhang, H., Cai, W., Effantin, G., Perot, J., Neyton, J., Chipot, C., Schoehn, G., Dehez, F., Nury, H.(2020) Structure 28: 1131-1140.e4
- PubMed: 32726573 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2020.07.004
- Primary Citation Related Structures:
6Y1Z - PubMed Abstract:
Inaccurately perceived as niche drugs, antiemetics are key elements of cancer treatment alleviating the most dreaded side effect of chemotherapy. Serotonin 5-HT3 receptor antagonists are the most commonly prescribed class of drugs to control chemotherapy-induced nausea and vomiting. These antagonists have been clinically successful drugs since the 1980s, yet our understanding of how they operate at the molecular level has been hampered by the difficulty of obtaining structures of drug-receptor complexes. Here, we report the cryoelectron microscopy structure of the palonosetron-bound 5-HT3 receptor. We investigate the binding of palonosetron, granisetron, dolasetron, ondansetron, and cilansetron using molecular dynamics, covering the whole set of antagonists used in clinical practice. The structural and computational results yield detailed atomic insight into the binding modes of the drugs. In light of our data, we establish a comprehensive framework underlying the inhibition mechanism by the -setron drug family.
- CNRS, Univ. Grenoble Alpes, CEA, IBS, 38000 Grenoble, France.
Organizational Affiliation: