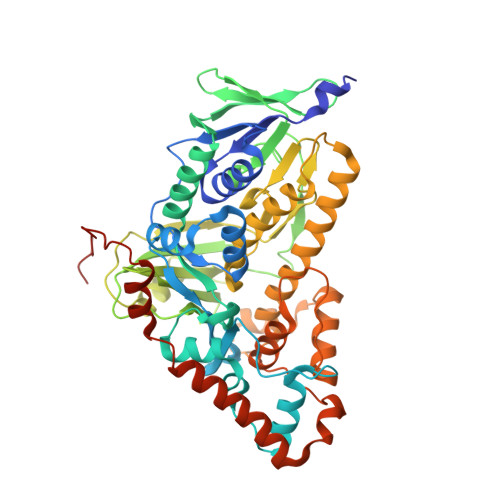
Structure of apo flavin-dependent halogenase Xcc4156 hints at a reason for cofactor-soaking difficulties.
Widmann, C., Ismail, M., Sewald, N., Niemann, H.H.(2020) Acta Crystallogr D Struct Biol 76: 687-697
- PubMed: 32627741 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798320007731
- Primary Citation Related Structures:
6Y1W - PubMed Abstract:
Flavin-dependent halogenases regioselectively introduce halide substituents into electron-rich substrates under mild reaction conditions. For the enzyme Xcc4156 from Xanthomonas campestris, the structure of a complex with the cofactor flavin adenine dinucleotide (FAD) and a bromide ion would be of particular interest as this enzyme exclusively brominates model substrates in vitro. Apo Xcc4156 crystals diffracted to 1.6 Å resolution. The structure revealed an open substrate-binding site lacking the loop regions that close off the active site and contribute to substrate binding in tryptophan halogenases. Therefore, Xcc4156 might accept larger substrates, possibly even peptides. Soaking of apo Xcc4156 crystals with FAD led to crumbling of the intergrown crystals. Around half of the crystals soaked with FAD did not diffract, while in the others there was no electron density for FAD. The FAD-binding loop, which changes its conformation between the apo and the FAD-bound form in related enzymes, is involved in a crystal contact in the apo Xcc4156 crystals. The conformational change that is predicted to occur upon FAD binding would disrupt this crystal contact, providing a likely explanation for the destruction of the apo crystals in the presence of FAD. Soaking with only bromide did not result in bromide bound to the catalytic halide-binding site. Simultaneous soaking with FAD and bromide damaged the crystals more severely than soaking with only FAD. Together, these latter two observations suggest that FAD and bromide bind to Xcc4156 with positive cooperativity. Thus, apo Xcc4156 crystals provide functional insight into FAD and bromide binding, even though neither the cofactor nor the halide is visible in the structure.
- Structural Biochemistry (BCIV), Department of Chemistry, Bielefeld University, Universitaetsstrasse 25, 33615 Bielefeld, Germany.
Organizational Affiliation: