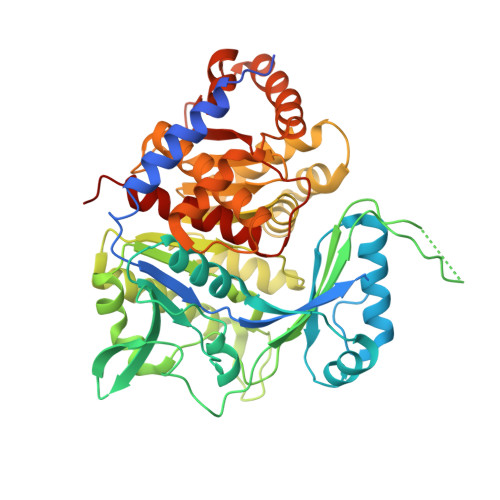
Structure of an ancestral ADP-dependent kinase with fructose-6P reveals key residues for binding, catalysis, and ligand-induced conformational changes.
Munoz, S.M., Castro-Fernandez, V., Guixe, V.(2020) J Biological Chem 296: 100219-100219
- PubMed: 33839685 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA120.015376
- Primary Citation Related Structures:
6XIO - PubMed Abstract:
ADP-dependent kinases were first described in archaea, although their presence has also been reported in bacteria and eukaryotes (human and mouse). This enzyme family comprises three substrate specificities; specific phosphofructokinases (ADP-PFKs), specific glucokinases (ADP-GKs), and bifunctional enzymes (ADP-PFK/GK). Although many structures are available for members of this family, none exhibits fructose-6-phosphate (F6P) at the active site. Using an ancestral enzyme, we obtain the first structure of an ADP-dependent kinase (AncMsPFK) with F6P at its active site. Key residues for sugar binding and catalysis were identified by alanine scanning, D36 being a critical residue for F6P binding and catalysis. However, this residue hinders glucose binding because its mutation to alanine converts the AncMsPFK enzyme into a specific ADP-GK. Residue K179 is critical for F6P binding, while residues N181 and R212 are also important for this sugar binding, but to a lesser extent. This structure also provides evidence for the requirement of both substrates (sugar and nucleotide) to accomplish the conformational change leading to a closed conformation. This suggests that AncMsPFK mainly populates two states (open and closed) during the catalytic cycle, as reported for specific ADP-PFK. This situation differs from that described for specific ADP-GK enzymes, where each substrate independently causes a sequential domain closure, resulting in three conformational states (open, semiclosed, and closed).
- Laboratorio de Bioquímica y Biología Molecular, Departamento de Biología, Facultad de Ciencias, Universidad de Chile, Santiago, Chile.
Organizational Affiliation: