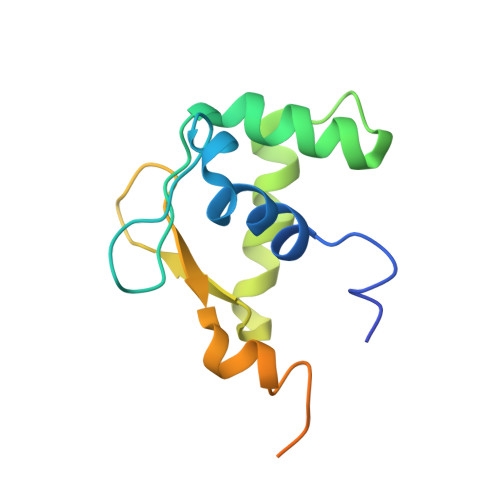
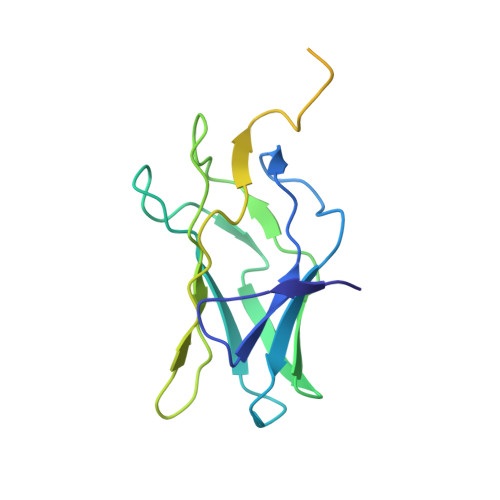
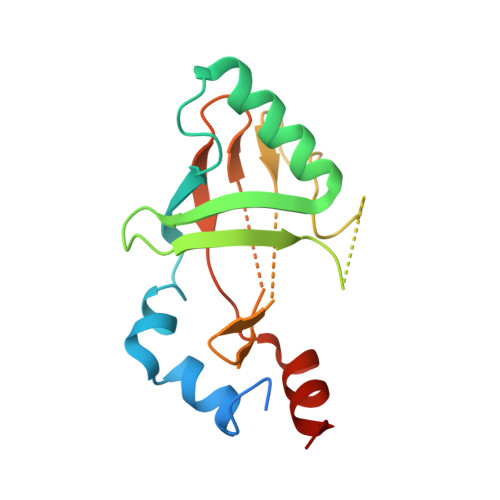
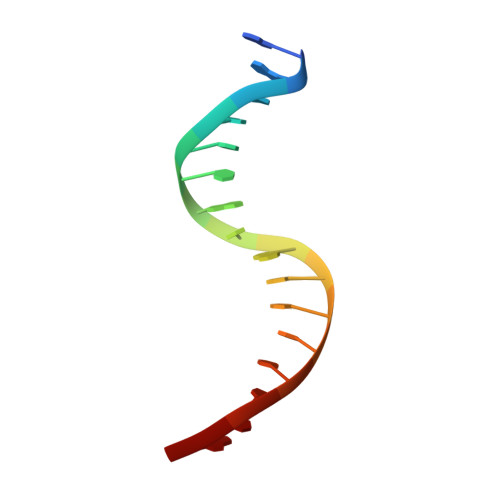
Allosteric interference in oncogenic FLI1 and ERG transactions by mithramycins.
Hou, C., Mandal, A., Rohr, J., Tsodikov, O.V.(2021) Structure 29: 404-412.e4
- PubMed: 33275876 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2020.11.012
- Primary Citation Related Structures:
6VG2, 6VG8, 6VGD, 6VGE, 6VGG - PubMed Abstract:
ETS family transcription factors of ERG and FLI1 play a key role in oncogenesis of prostate cancer and Ewing sarcoma by binding regulatory DNA sites and interfering with function of other factors. Mithramycin (MTM) is an anti-cancer, DNA binding natural product that functions as a potent antagonist of ERG and FLI1 by an unknown mechanism. We present a series of crystal structures of the DNA binding domain (DBD) of ERG/FLI1 culminating in a structure of a high-order complex of the ERG/FLI1 DBD, transcription factor Runx2, core-binding factor beta (Cbfβ), and MTM on a DNA enhancer site, along with supporting DNA binding studies using MTM and its analogues. Taken together, these data provide insight into allosteric mechanisms underlying ERG and FLI1 transactions and their disruption by MTM analogues.
- Department of Pharmaceutical Sciences, College of Pharmacy, University of Kentucky, Lexington, KY 40536, USA.
Organizational Affiliation: