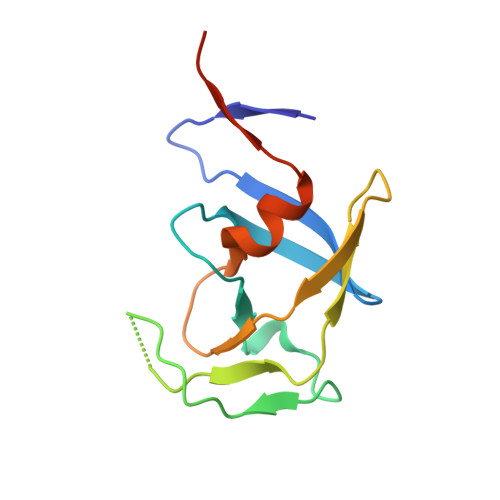
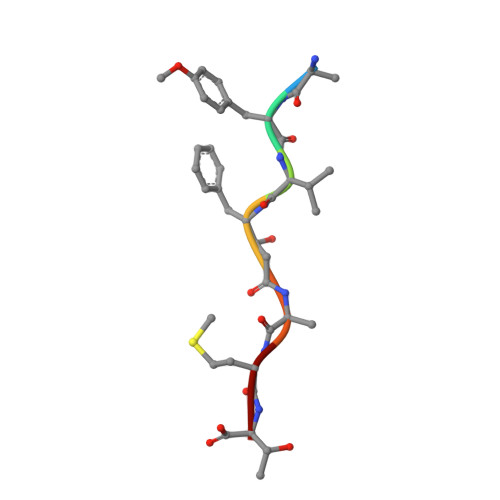
Comparison of a retroviral protease in monomeric and dimeric states.
Wosicki, S., Gilski, M., Zabranska, H., Pichova, I., Jaskolski, M.(2019) Acta Crystallogr D Struct Biol 75: 904-917
- PubMed: 31588922 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798319011355
- Primary Citation Related Structures:
6S1U, 6S1V, 6S1W - PubMed Abstract:
Retroviral proteases (RPs) are of high interest owing to their crucial role in the maturation process of retroviral particles. RPs are obligatory homodimers, with a pepsin-like active site built around two aspartates (in DTG triads) that activate a water molecule, as the nucleophile, under two flap loops. Mason-Pfizer monkey virus (M-PMV) is unique among retroviruses as its protease is also stable in the monomeric form, as confirmed by an existing crystal structure of a 13 kDa variant of the protein (M-PMV PR) and its previous biochemical characterization. In the present work, two mutants of M-PMV PR, D26N and C7A/D26N/C106A, were crystallized in complex with a peptidomimetic inhibitor and one mutant (D26N) was crystallized without the inhibitor. The crystal structures were solved at resolutions of 1.6, 1.9 and 2.0 Å, respectively. At variance with the previous study, all of the new structures have the canonical dimeric form of retroviral proteases. The protomers within a dimer differ mainly in the flap-loop region, with the most extreme case observed in the apo structure, in which one flap loop is well defined while the other flap loop is not defined by electron density. The presence of the inhibitor molecules in the complex structures was assessed using polder maps, but some details of their conformations remain ambiguous. In all of the presented structures the active site contains a water molecule buried deeply between the Asn26-Thr27-Gly28 triads of the protomers. Such a water molecule is completely unique not only in retropepsins but also in aspartic proteases in general. The C7A and C106A mutations do not influence the conformation of the protein. The Cys106 residue is properly placed at the homodimer interface area for a disulfide cross-link, but the reducing conditions of the crystallization experiment prevented S-S bond formation. An animated Interactive 3D Complement (I3DC) is available in Proteopedia at http://proteopedia.org/w/Journal:Acta_Cryst_D:S2059798319011355.
- Center for Biocrystallographic Research, Institute of Bioorganic Chemistry, Polish Academy of Sciences, 61-704 Poznan, Poland.
Organizational Affiliation: