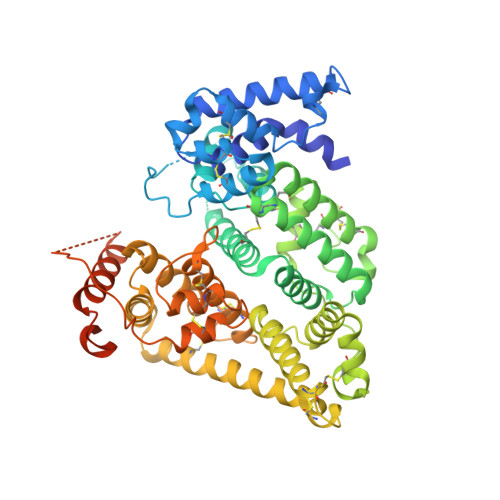
Controlled dehydration, structural flexibility and gadolinium MRI contrast compound binding in the human plasma glycoprotein afamin.
Naschberger, A., Juyoux, P., von Velsen, J., Rupp, B., Bowler, M.W.(2019) Acta Crystallogr D Struct Biol 75: 1071-1083
- PubMed: 31793901 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798319013500
- Primary Citation Related Structures:
6FAK, 6RQ7 - PubMed Abstract:
Afamin, which is a human blood plasma glycoprotein, a putative multifunctional transporter of hydrophobic molecules and a marker for metabolic syndrome, poses multiple challenges for crystallographic structure determination, both practically and in analysis of the models. Several hundred crystals were analysed, and an unusual variability in cell volume and difficulty in solving the structure despite an ∼34% sequence identity with nonglycosylated human serum albumin indicated that the molecule exhibits variable and context-sensitive packing, despite the simplified glycosylation in insect cell-expressed recombinant afamin. Controlled dehydration of the crystals was able to stabilize the orthorhombic crystal form, reducing the number of molecules in the asymmetric unit from the monoclinic form and changing the conformational state of the protein. An iterative strategy using fully automatic experiments available on MASSIF-1 was used to quickly determine the optimal protocol to achieve the phase transition, which should be readily applicable to many types of sample. The study also highlights the drawback of using a single crystallographic structure model for computational modelling purposes given that the conformational state of the binding sites and the electron density in the binding site, which is likely to result from PEGs, greatly varies between models. This also holds for the analysis of nonspecific low-affinity ligands, where often a variety of fragments with similar uncertainty can be modelled, inviting interpretative bias. As a promiscuous transporter, afamin also seems to bind gadoteridol, a magnetic resonance imaging contrast compound, in at least two sites. One pair of gadoteridol molecules is located near the human albumin Sudlow site, and a second gadoteridol molecule is located at an intermolecular site in proximity to domain IA. The data from the co-crystals support modern metrics of data quality in the context of the information that can be gleaned from data sets that would be abandoned on classical measures.
- Department of Genetic Epidemiology, Medical University Innsbruck, Schöpfstrasse 41, A-6020 Innsbruck, Austria.
Organizational Affiliation: