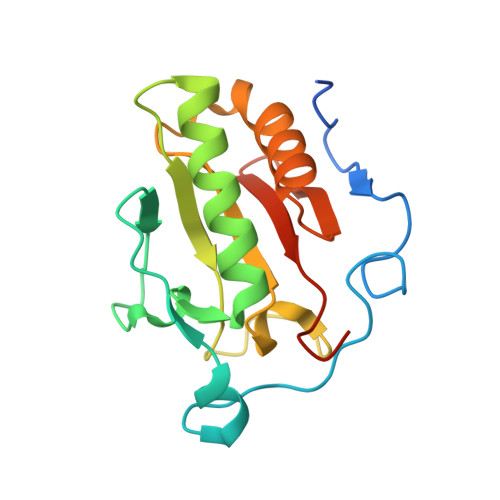
Structure of Sonic Hedgehog protein in complex with zinc(II) and magnesium(II) reveals ion-coordination plasticity relevant to peptide drug design.
Bonn-Breach, R., Gu, Y., Jenkins, J., Fasan, R., Wedekind, J.(2019) Acta Crystallogr D Struct Biol 75: 969-979
- PubMed: 31692471 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798319012890
- Primary Citation Related Structures:
6PJV - PubMed Abstract:
The Hedgehog pathway is an essential cell-signaling paradigm implicated in cancer tumorigenesis and the developmental disorder holoprosencephaly, making it an attractive target for therapeutic design. The N-terminal domain of the Sonic Hedgehog protein (Shh-N) is the essential signaling molecule in the Hedgehog pathway. In this role Shh-N interacts with its cognate membrane receptor Patched, as well as the regulatory proteins HHIP and CDO, by utilizing interfaces harboring one or more divalent ions. Here, the crystal structure of human Shh-N is presented at 1.43 Å resolution, representing a landmark in the characterization of this protein. The structure reveals that the conserved Zn 2+ -binding site adopts an atypical octahedral coordination geometry, whereas an adjacent binding site, normally occupied by binuclear Ca 2+ , has been supplanted by a single octahedrally bound Mg 2+ . Both divalent sites are compared with those in previous Shh-N structures, which demonstrates a significant degree of plasticity of the Shh-N protein in terms of divalent ion binding. The presence of a high Mg 2+ concentration in the crystallization medium appears to have influenced metal loading at both metal ion-binding sites. These observations have technical and design implications for efforts focused on the development of inhibitors that target Shh-N-mediated protein-protein interactions.
- Department of Biochemistry and Biophysics, University of Rochester School of Medicine and Dentistry, 601 Elmwood Avenue, Rochester, NY 14642, USA.
Organizational Affiliation: