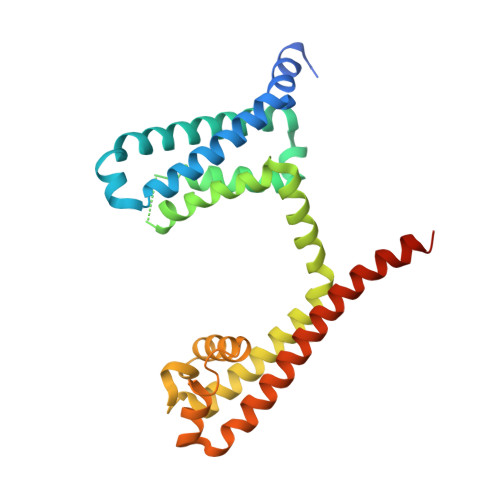
Resting-State Structure and Gating Mechanism of a Voltage-Gated Sodium Channel.
Wisedchaisri, G., Tonggu, L., McCord, E., Gamal El-Din, T.M., Wang, L., Zheng, N., Catterall, W.A.(2019) Cell 178: 993
- PubMed: 31353218 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2019.06.031
- Primary Citation Related Structures:
6P6W, 6P6X, 6P6Y - PubMed Abstract:
Voltage-gated sodium (Na V ) channels initiate action potentials in nerve, muscle, and other electrically excitable cells. The structural basis of voltage gating is uncertain because the resting state exists only at deeply negative membrane potentials. To stabilize the resting conformation, we inserted voltage-shifting mutations and introduced a disulfide crosslink in the VS of the ancestral bacterial sodium channel Na V Ab. Here, we present a cryo-EM structure of the resting state and a complete voltage-dependent gating mechanism. The S4 segment of the VS is drawn intracellularly, with three gating charges passing through the transmembrane electric field. This movement forms an elbow connecting S4 to the S4-S5 linker, tightens the collar around the S6 activation gate, and prevents its opening. Our structure supports the classical "sliding helix" mechanism of voltage sensing and provides a complete gating mechanism for voltage sensor function, pore opening, and activation-gate closure based on high-resolution structures of a single sodium channel protein.
- Department of Pharmacology, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: