Structure Based Prediction of Neoantigen Immunogenicity.
Riley, T.P., Keller, G.L.J., Smith, A.R., Davancaze, L.M., Arbuiso, A.G., Devlin, J.R., Baker, B.M.(2019) Front Immunol 10: 2047-2047
- PubMed: 31555277 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fimmu.2019.02047
- Primary Citation Related Structures:
6OPD, 6PTB, 6PTE - PubMed Abstract:
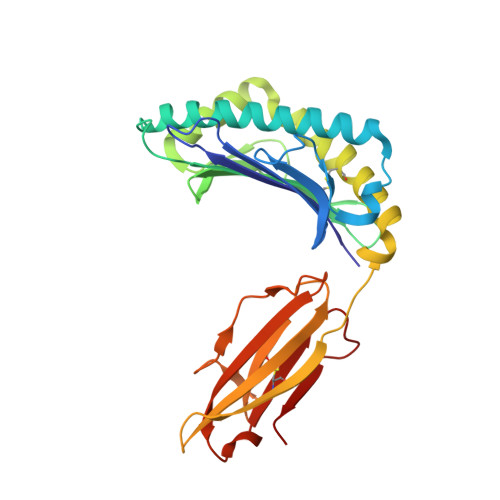
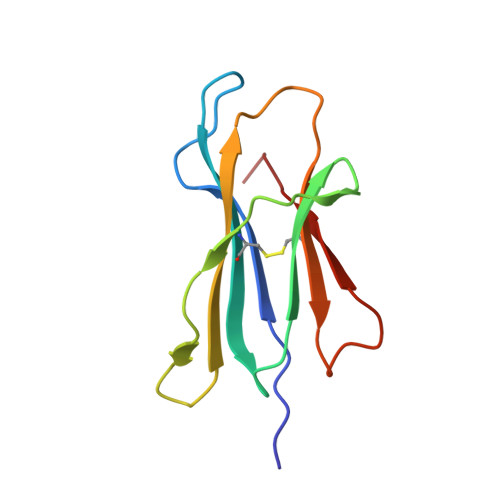
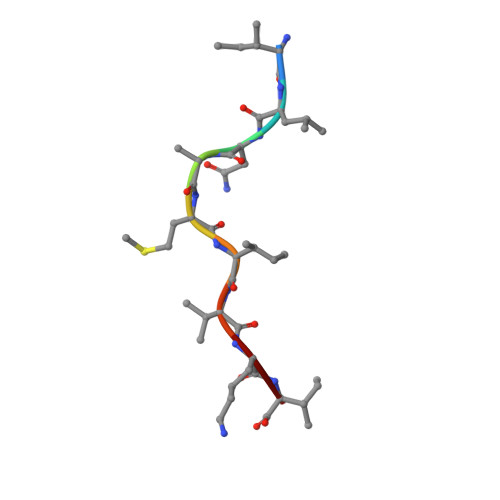
The development of immunological therapies that incorporate peptide antigens presented to T cells by MHC proteins is a long sought-after goal, particularly for cancer, where mutated neoantigens are being explored as personalized cancer vaccines. Although neoantigens can be identified through sequencing, bioinformatics and mass spectrometry, identifying those which are immunogenic and able to promote tumor rejection remains a significant challenge. Here we examined the potential of high-resolution structural modeling followed by energetic scoring of structural features for predicting neoantigen immunogenicity. After developing a strategy to rapidly and accurately model nonameric peptides bound to the common class I MHC protein HLA-A2, we trained a neural network on structural features that influence T cell receptor (TCR) and peptide binding energies. The resulting structurally-parameterized neural network outperformed methods that do not incorporate explicit structural or energetic properties in predicting CD8 + T cell responses of HLA-A2 presented nonameric peptides, while also providing insight into the underlying structural and biophysical mechanisms governing immunogenicity. Our proof-of-concept study demonstrates the potential for structure-based immunogenicity predictions in the development of personalized peptide-based vaccines.
- Department of Chemistry and Biochemistry and the Harper Cancer Research Institute, University of Notre Dame, Notre Dame, IN, United States.
Organizational Affiliation: