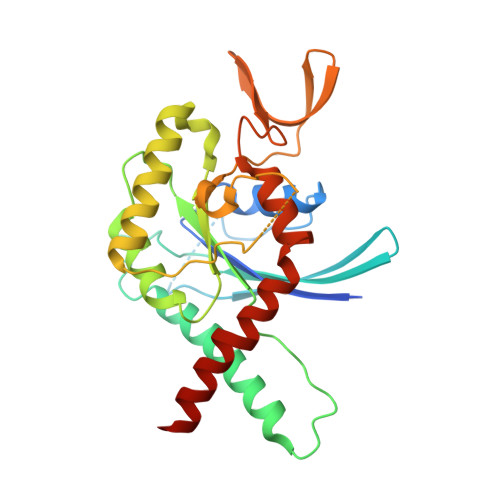
Revisiting SEPT7 and the slippage of beta-strands in the septin family.
Brognara, G., Pereira, H.M., Brandao-Neto, J., Araujo, A.P.U., Garratt, R.C.(2019) J Struct Biol 207: 67-73
- PubMed: 31009756 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2019.04.015
- Primary Citation Related Structures:
6N0B, 6N12 - PubMed Abstract:
Septins are GTP-binding proteins that will often spontaneously assemble into filaments. In some species, particularly budding yeast, it is well known that these are capable of associating with membranes in order to fulfill their cellular role as a component of the cytoskeleton. Different from other human septins, SEPT7 appears to be unique in that it is an essential component of all hetero-oligomeric complexes described to date. As a step towards understanding the molecular basis of filament assembly, here we present two high-resolution structures of the SEPT7 GTPase domain complexed with GDP. One of these reveals a previously unreported coordination for the magnesium ion involving four water molecules and only a tenuous connection to the protein. The higher resolution structures provide unambiguous insight into the interactions at the G-interface where a structural motif based on an antiparallel β-bridge allows for the rationalization of why some septins show nucleotide-dependent β-strand slippage and others do not.
- São Carlos Institute of Physics, University of São Paulo, São Carlos, São Paulo, Brazil.
Organizational Affiliation: