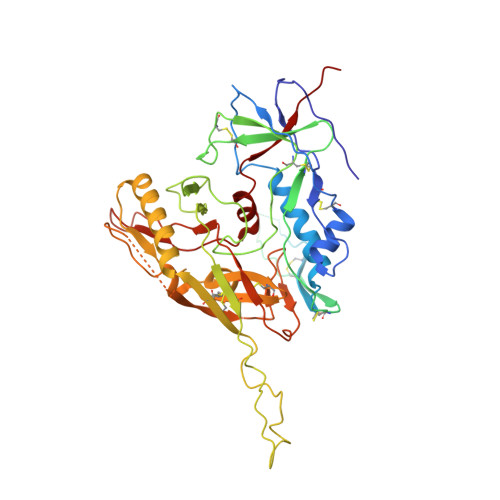
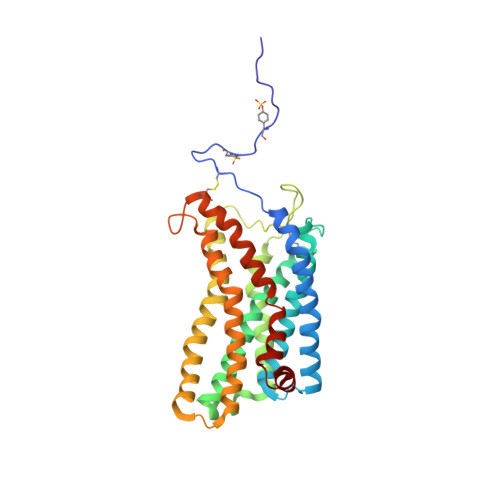
Structural basis of coreceptor recognition by HIV-1 envelope spike.
Shaik, M.M., Peng, H., Lu, J., Rits-Volloch, S., Xu, C., Liao, M., Chen, B.(2018) Nature 565: 318-323
- PubMed: 30542158 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-018-0804-9
- Primary Citation Related Structures:
6MEO, 6MET - PubMed Abstract:
HIV-1 envelope glycoprotein (Env), which consists of trimeric (gp160) 3 cleaved to (gp120 and gp41) 3 , interacts with the primary receptor CD4 and a coreceptor (such as chemokine receptor CCR5) to fuse viral and target-cell membranes. The gp120-coreceptor interaction has previously been proposed as the most crucial trigger for unleashing the fusogenic potential of gp41. Here we report a cryo-electron microscopy structure of a full-length gp120 in complex with soluble CD4 and unmodified human CCR5, at 3.9 Å resolution. The V3 loop of gp120 inserts into the chemokine-binding pocket formed by seven transmembrane helices of CCR5, and the N terminus of CCR5 contacts the CD4-induced bridging sheet of gp120. CCR5 induces no obvious allosteric changes in gp120 that can propagate to gp41; it does bring the Env trimer close to the target membrane. The N terminus of gp120, which is gripped by gp41 in the pre-fusion or CD4-bound Env, flips back in the CCR5-bound conformation and may irreversibly destabilize gp41 to initiate fusion. The coreceptor probably functions by stabilizing and anchoring the CD4-induced conformation of Env near the cell membrane. These results advance our understanding of HIV-1 entry into host cells and may guide the development of vaccines and therapeutic agents.
- Division of Molecular Medicine, Boston Children's Hospital, Harvard Medical School, Boston, MA, USA.
Organizational Affiliation: