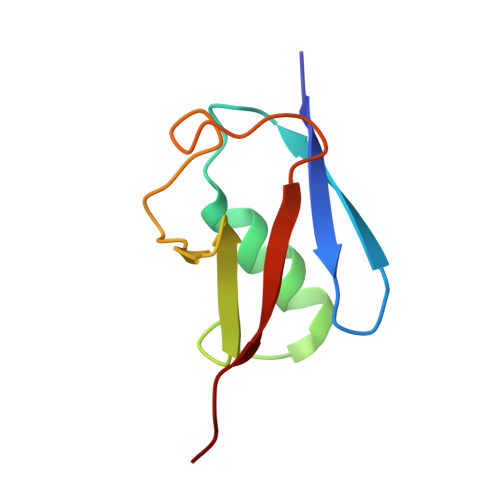
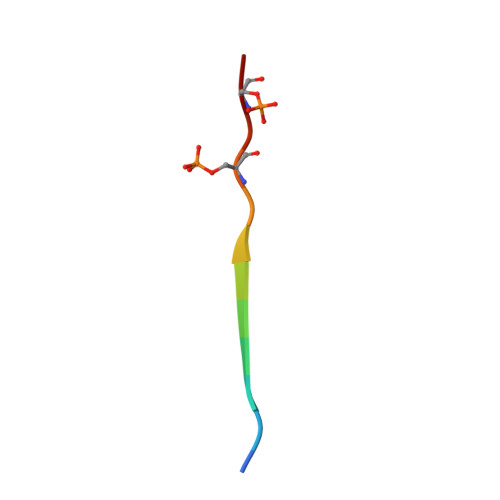
Casein kinase-2-mediated phosphorylation increases the SUMO-dependent activity of the cytomegalovirus transactivator IE2.
Tripathi, V., Chatterjee, K.S., Das, R.(2019) J Biological Chem 294: 14546-14561
- PubMed: 31371453 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.009601
- Primary Citation Related Structures:
6K5R, 6K5T - PubMed Abstract:
Many viral factors manipulate the host post-translational modification (PTM) machinery for efficient viral replication. In particular, phosphorylation and SUMOylation can distinctly regulate the activity of the human cytomegalovirus (HCMV) transactivator immediate early 2 (IE2). However, the molecular mechanism of this process is unknown. Using various structural, biochemical, and cell-based approaches, here we uncovered that IE2 exploits a cross-talk between phosphorylation and SUMOylation. A scan for small ubiquitin-like modifier (SUMO)-interacting motifs (SIMs) revealed two SIMs in IE2, and a real-time SUMOylation assay indicated that the N-terminal SIM (IE2-SIM1) enhances IE2 SUMOylation up to 4-fold. Kinetic analysis and structural studies disclosed that IE2 is a SUMO cis- E3 ligase. We also found that two putative casein kinase 2 (CK2) sites adjacent to IE2-SIM1 are phosphorylated in vitro and in cells. The phosphorylation drastically increased IE2-SUMO affinity, IE2 SUMOylation, and cis- E3 activity of IE2. Additional salt bridges between the phosphoserines and SUMO accounted for the increased IE2-SUMO affinity. Phosphorylation also enhanced the SUMO-dependent transactivation activity and auto-repression activity of IE2. Together, our findings highlight a novel mechanism whereby SUMOylation and phosphorylation of the viral cis- E3 ligase and transactivator protein IE2 work in tandem to enable transcriptional regulation of viral gene.
- National Centre for Biological Sciences, Tata Institute of Fundamental Research, Bengaluru-560065, India.
Organizational Affiliation: