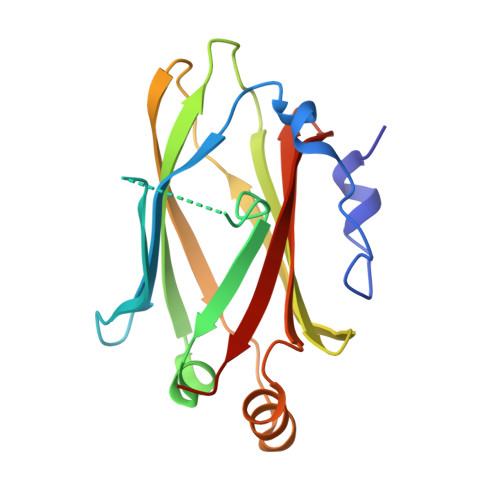
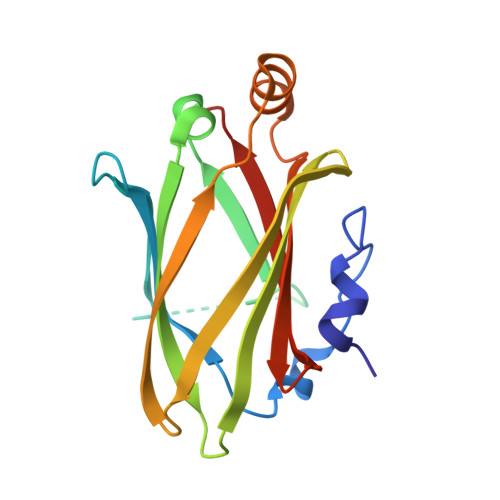
The Ciliary Machinery Is Repurposed for T Cell Immune Synapse Trafficking of LCK.
Stephen, L.A., ElMaghloob, Y., McIlwraith, M.J., Yelland, T., Castro Sanchez, P., Roda-Navarro, P., Ismail, S.(2018) Dev Cell 47: 122-132.e4
- PubMed: 30220567 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.devcel.2018.08.012
- Primary Citation Related Structures:
6H6A - PubMed Abstract:
Upon engagement of the T cell receptor with an antigen-presenting cell, LCK initiates TCR signaling by phosphorylating its activation motifs. However, the mechanism of LCK activation specifically at the immune synapse is a major question. We show that phosphorylation of the LCK activating Y394, despite modestly increasing its catalytic rate, dramatically focuses LCK localization to the immune synapse. We describe a trafficking mechanism whereby UNC119A extracts membrane-bound LCK by sequestering the hydrophobic myristoyl group, followed by release at the target membrane under the control of the ciliary ARL3/ARL13B. The UNC119A N terminus acts as a "regulatory arm" by binding the LCK kinase domain, an interaction inhibited by LCK Y394 phosphorylation, thus together with the ARL3/ARL13B machinery ensuring immune synapse focusing of active LCK. We propose that the ciliary machinery has been repurposed by T cells to generate and maintain polarized segregation of signals such as activated LCK at the immune synapse.
- CR-UK Beatson Institute, Garscube Estate, Switchback Road, Glasgow G61 1BD, UK.
Organizational Affiliation: