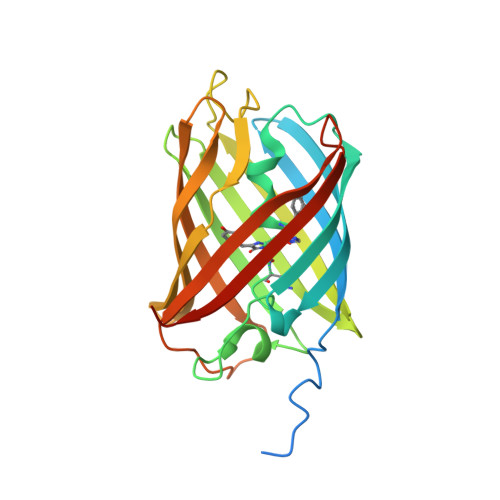
Mechanistic investigation of mEos4b reveals a strategy to reduce track interruptions in sptPALM.
De Zitter, E., Thedie, D., Monkemoller, V., Hugelier, S., Beaudouin, J., Adam, V., Byrdin, M., Van Meervelt, L., Dedecker, P., Bourgeois, D.(2019) Nat Methods 16: 707-710
- PubMed: 31285624 Search on PubMed
- DOI: https://doi.org/10.1038/s41592-019-0462-3
- Primary Citation Related Structures:
6GOY, 6GP0, 6GP1 - PubMed Abstract:
Green-to-red photoconvertible fluorescent proteins repeatedly enter dark states, causing interrupted tracks in single-particle-tracking localization microscopy (sptPALM). We identified a long-lived dark state in photoconverted mEos4b that results from isomerization of the chromophore and efficiently absorbs cyan light. Addition of weak 488-nm light swiftly reverts this dark state to the fluorescent state. This strategy largely eliminates slow blinking and enables the recording of longer tracks in sptPALM with minimum effort.
- Department of Chemistry, KU Leuven, Heverlee, Belgium.
Organizational Affiliation: