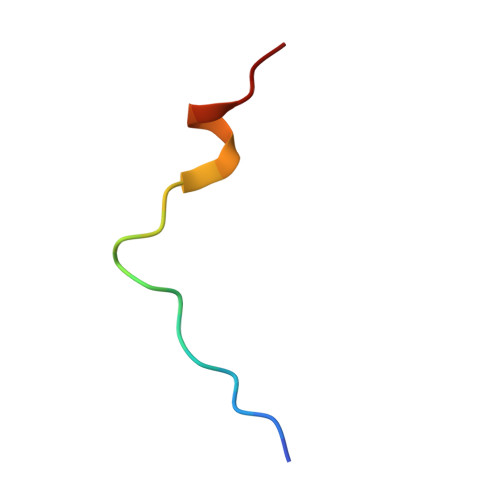
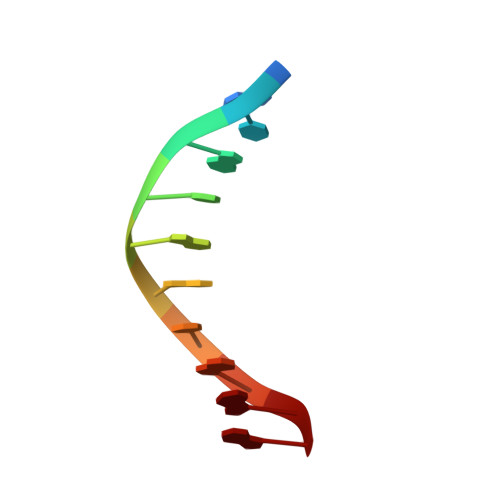
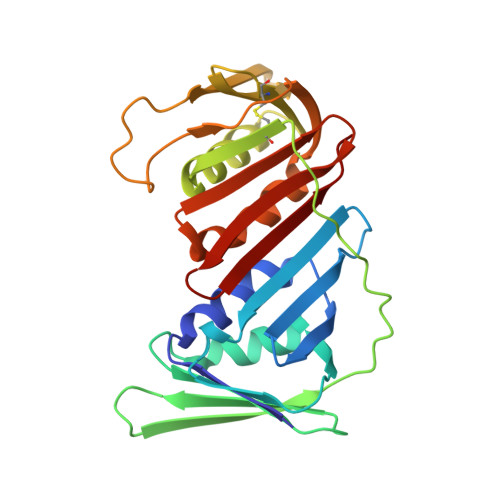
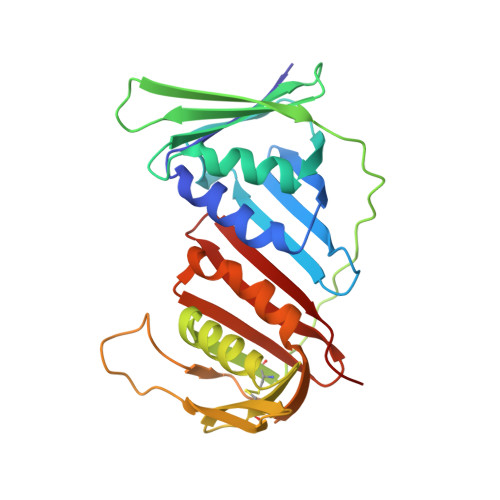
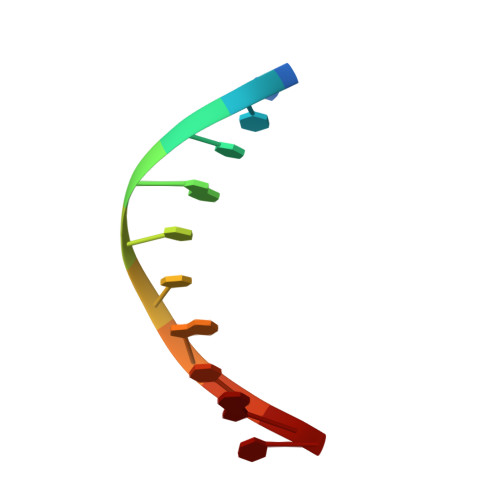
p15PAF binding to PCNA modulates the DNA sliding surface.
De March, M., Barrera-Vilarmau, S., Crespan, E., Mentegari, E., Merino, N., Gonzalez-Magana, A., Romano-Moreno, M., Maga, G., Crehuet, R., Onesti, S., Blanco, F.J., De Biasio, A.(2018) Nucleic Acids Res 46: 9816-9828
- PubMed: 30102405 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gky723
- Primary Citation Related Structures:
6EHT, 6GWS - PubMed Abstract:
p15PAF is an oncogenic intrinsically disordered protein that regulates DNA replication and lesion bypass by interacting with the human sliding clamp PCNA. In the absence of DNA, p15PAF traverses the PCNA ring via an extended PIP-box that contacts the sliding surface. Here, we probed the atomic-scale structure of p15PAF-PCNA-DNA ternary complexes. Crystallography and MD simulations show that, when p15PAF occupies two subunits of the PCNA homotrimer, DNA within the ring channel binds the unoccupied subunit. The structure of PCNA-bound p15PAF in the absence and presence of DNA is invariant, and solution NMR confirms that DNA does not displace p15PAF from the ring wall. Thus, p15PAF reduces the available sliding surfaces of PCNA, and may function as a belt that fastens the DNA to the clamp during synthesis by the replicative polymerase (pol δ). This constraint, however, may need to be released for efficient DNA lesion bypass by the translesion synthesis polymerase (pol η). Accordingly, our biochemical data show that p15PAF impairs primer synthesis by pol η-PCNA holoenzyme against both damaged and normal DNA templates. In light of our findings, we discuss the possible mechanistic roles of p15PAF in DNA replication and suppression of DNA lesion bypass.
- Structural Biology Laboratory, Elettra-Sincrotrone Trieste S.C.p.A., Trieste 34149, Italy.
Organizational Affiliation: