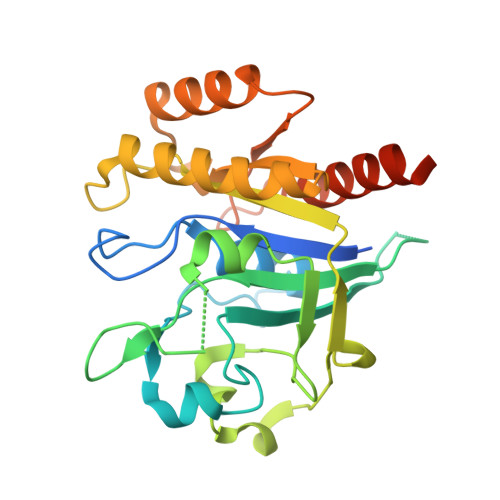
The N-Terminal GTPase Domain of p190RhoGAP Proteins Is a PseudoGTPase.
Stiegler, A.L., Boggon, T.J.(2018) Structure 26: 1451
- PubMed: 30174148 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2018.07.015
- Primary Citation Related Structures:
6D4G - PubMed Abstract:
The pseudoGTPases are a rapidly growing and important group of pseudoenzymes. p190RhoGAP proteins are critical regulators of Rho signaling and contain two previously identified pseudoGTPase domains. Here we report that p190RhoGAP proteins contain a third pseudoGTPase domain, termed N-GTPase. We find that GTP constitutively purifies with the N-GTPase domain, and a 2.8-Å crystal structure of p190RhoGAP-A co-purified with GTP reveals an unusual GTP-Mg 2+ binding pocket. Six inserts in N-GTPase indicate perturbed catalytic activity and inability to bind to canonical GTPase activating proteins, guanine nucleotide exchange factors, and effector proteins. Biochemical analysis shows that N-GTPase does not detectably hydrolyze GTP, and exchanges nucleotide only under harsh Mg 2+ chelation. Furthermore, mutational analysis shows that GTP and Mg 2+ binding stabilizes the domain. Therefore, our results support that N-GTPase is a nucleotide binding, non-hydrolyzing, pseudoGTPase domain that may act as a protein-protein interaction domain. Thus, unique among known proteins, p190RhoGAPs contain three pseudoGTPase domains.
- Department of Pharmacology, Yale University School of Medicine, 333 Cedar Street, New Haven, CT 06520, USA.
Organizational Affiliation: