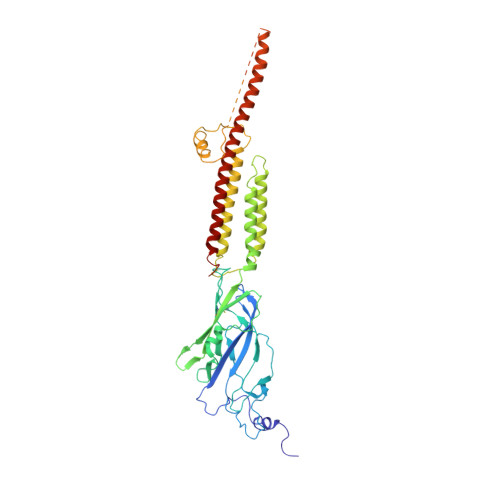
Cryo-EM structure of 5-HT
Basak, S., Gicheru, Y., Samanta, A., Molugu, S.K., Huang, W., Fuente, M., Hughes, T., Taylor, D.J., Nieman, M.T., Moiseenkova-Bell, V., Chakrapani, S.(2018) Nat Commun 9: 514-514
- PubMed: 29410406 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-02997-4
- Primary Citation Related Structures:
6BE1 - PubMed Abstract:
Serotonin receptors (5-HT 3A R) directly regulate gut movement, and drugs that inhibit 5-HT 3A R function are used to control emetic reflexes associated with gastrointestinal pathologies and cancer therapies. The 5-HT 3A R function involves a finely tuned orchestration of three domain movements that include the ligand-binding domain, the pore domain, and the intracellular domain. Here, we present the structure from the full-length 5-HT 3A R channel in the apo-state determined by single-particle cryo-electron microscopy at a nominal resolution of 4.3 Å. In this conformation, the ligand-binding domain adopts a conformation reminiscent of the unliganded state with the pore domain captured in a closed conformation. In comparison to the 5-HT 3A R crystal structure, the full-length channel in the apo-conformation adopts a more expanded conformation of all the three domains with a characteristic twist that is implicated in gating.
- Department of Physiology and Biophysics, Case Western Reserve University, Cleveland, OH, 44106-4970, USA.
Organizational Affiliation: