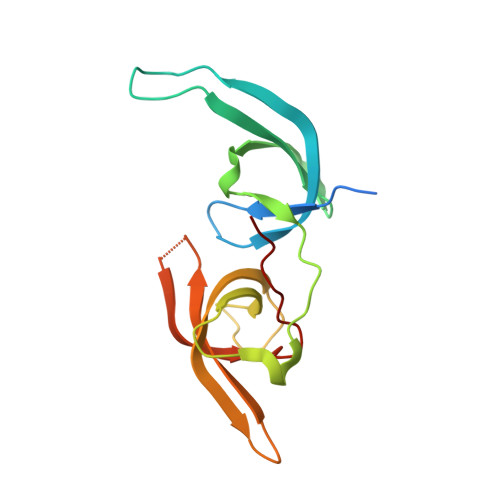
Structure of the UHRF1 Tandem Tudor Domain Bound to a Methylated Non-histone Protein, LIG1, Reveals Rules for Binding and Regulation.
Kori, S., Ferry, L., Matano, S., Jimenji, T., Kodera, N., Tsusaka, T., Matsumura, R., Oda, T., Sato, M., Dohmae, N., Ando, T., Shinkai, Y., Defossez, P.A., Arita, K.(2019) Structure 27: 485
- PubMed: 30639225 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2018.11.012
- Primary Citation Related Structures:
5YY9, 5YYA - PubMed Abstract:
The protein UHRF1 is crucial for DNA methylation maintenance. The tandem Tudor domain (TTD) of UHRF1 binds histone H3K9me2/3 with micromolar affinity, as well as unmethylated linker regions within UHRF1 itself, causing auto-inhibition. Recently, we showed that a methylated histone-like region of DNA ligase 1 (LIG1K126me2/me3) binds the UHRF1 TTD with nanomolar affinity, permitting UHRF1 recruitment to chromatin. Here we report the crystal structure of the UHRF1 TTD bound to a LIG1K126me3 peptide. The data explain the basis for the high TTD-binding affinity of LIG1K126me3 and reveal that the interaction may be regulated by phosphorylation. Binding of LIG1K126me3 switches the overall structure of UHRF1 from a closed to a flexible conformation, suggesting that auto-inhibition is relieved. Our results provide structural insight into how UHRF1 performs its key function in epigenetic maintenance.
- Graduate School of Medical Life Science, Yokohama City University, Yokohama 230-0045, Japan.
Organizational Affiliation: