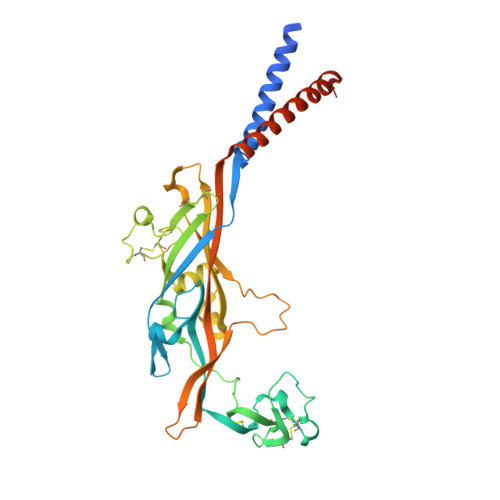
Druggable negative allosteric site of P2X3 receptors.
Wang, J., Wang, Y., Cui, W.W., Huang, Y., Yang, Y., Liu, Y., Zhao, W.S., Cheng, X.Y., Sun, W.S., Cao, P., Zhu, M.X., Wang, R., Hattori, M., Yu, Y.(2018) Proc Natl Acad Sci U S A 115: 4939-4944
- PubMed: 29674445 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1800907115
- Primary Citation Related Structures:
5YVE - PubMed Abstract:
Allosteric modulation provides exciting opportunities for drug discovery of enzymes, ion channels, and G protein-coupled receptors. As cation channels gated by extracellular ATP, P2X receptors have attracted wide attention as new drug targets. Although small molecules targeting P2X receptors have entered into clinical trials for rheumatoid arthritis, cough, and pain, negative allosteric modulation of these receptors remains largely unexplored. Here, combining X-ray crystallography, computational modeling, and functional studies of channel mutants, we identified a negative allosteric site on P2X3 receptors, fostered by the left flipper (LF), lower body (LB), and dorsal fin (DF) domains. Using two structurally analogous subtype-specific allosteric inhibitors of P2X3, AF-353 and AF-219, the latter being a drug candidate under phase II clinical trials for refractory chronic cough and idiopathic pulmonary fibrosis, we defined the molecular interactions between the drugs and receptors and the mechanism by which allosteric changes in the LF, DF, and LB domains modulate ATP activation of P2X3. Our detailed characterization of this druggable allosteric site should inspire new strategies to develop P2X3-specific allosteric modulators for clinical use.
- State Key Laboratory of Genetic Engineering, Collaborative Innovation Center of Genetics and Development, Department of Physiology and Biophysics, School of Life Sciences, Fudan University, 200438 Shanghai, China.
Organizational Affiliation: